Probe CUST_6425_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6425_PI426222305 | JHI_St_60k_v1 | DMT400014363 | ACTGGCCCGATTTCTCGGTATTTTATTCTTCTTCCTCCACAATTATTGTTCGTGTTGTTG |
All Microarray Probes Designed to Gene DMG400005633
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6622_PI426222305 | JHI_St_60k_v1 | DMT400014362 | AATCAATCTTCGCCATGGACTAATGGAATCCGACCCCATTTCAATCCTGATGACTACTGG |
| CUST_6425_PI426222305 | JHI_St_60k_v1 | DMT400014363 | ACTGGCCCGATTTCTCGGTATTTTATTCTTCTTCCTCCACAATTATTGTTCGTGTTGTTG |
Microarray Signals from CUST_6425_PI426222305
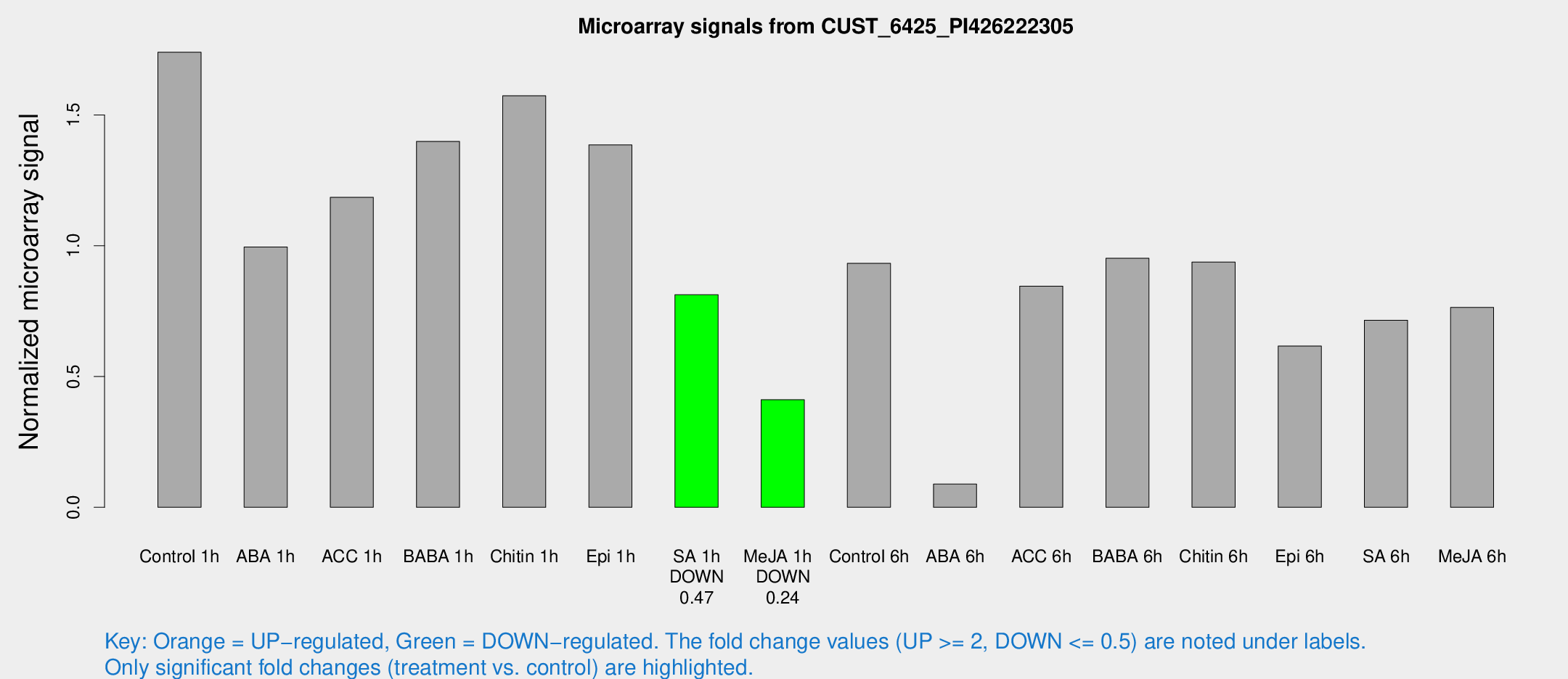
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 413.314 | 89.2452 | 1.74008 | 0.362975 |
| ABA 1h | 207.731 | 45.2147 | 0.99542 | 0.158118 |
| ACC 1h | 280.766 | 47.4393 | 1.18509 | 0.129475 |
| BABA 1h | 315.854 | 59.5461 | 1.39911 | 0.152512 |
| Chitin 1h | 331.252 | 64.1658 | 1.57351 | 0.214073 |
| Epi 1h | 269.524 | 15.8993 | 1.38579 | 0.0816635 |
| SA 1h | 194.757 | 34.9693 | 0.812926 | 0.126477 |
| Me-JA 1h | 75.635 | 5.46986 | 0.411112 | 0.0296642 |
| Control 6h | 227.747 | 58.0449 | 0.932739 | 0.205376 |
| ABA 6h | 22.0865 | 4.30046 | 0.0888339 | 0.0213047 |
| ACC 6h | 225.426 | 40.2184 | 0.845797 | 0.0840209 |
| BABA 6h | 257.086 | 66.0511 | 0.952205 | 0.244653 |
| Chitin 6h | 225.145 | 14.5776 | 0.938021 | 0.065726 |
| Epi 6h | 165.137 | 40.1782 | 0.616404 | 0.171362 |
| SA 6h | 160.343 | 15.376 | 0.714987 | 0.145189 |
| Me-JA 6h | 199.495 | 68.0314 | 0.764216 | 0.281748 |
Source Transcript PGSC0003DMT400014363 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G68520.1 | +3 | 5e-57 | 198 | 125/314 (40%) | B-box type zinc finger protein with CCT domain | chr1:25709331-25710749 REVERSE LENGTH=406 |
