Probe CUST_5768_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5768_PI426222305 | JHI_St_60k_v1 | DMT400023124 | CCTTTGAGTTATAGTAATGAAAATTGTGCAAGCTTGGCTTAAAGGTTTTGAACAGACGAA |
All Microarray Probes Designed to Gene DMG400008954
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5481_PI426222305 | JHI_St_60k_v1 | DMT400023125 | CAACCCACCCCTTAATTAAATGGAAGGCAAATAAATGTGTACAAATATTCCCCCATAATA |
| CUST_5768_PI426222305 | JHI_St_60k_v1 | DMT400023124 | CCTTTGAGTTATAGTAATGAAAATTGTGCAAGCTTGGCTTAAAGGTTTTGAACAGACGAA |
Microarray Signals from CUST_5768_PI426222305
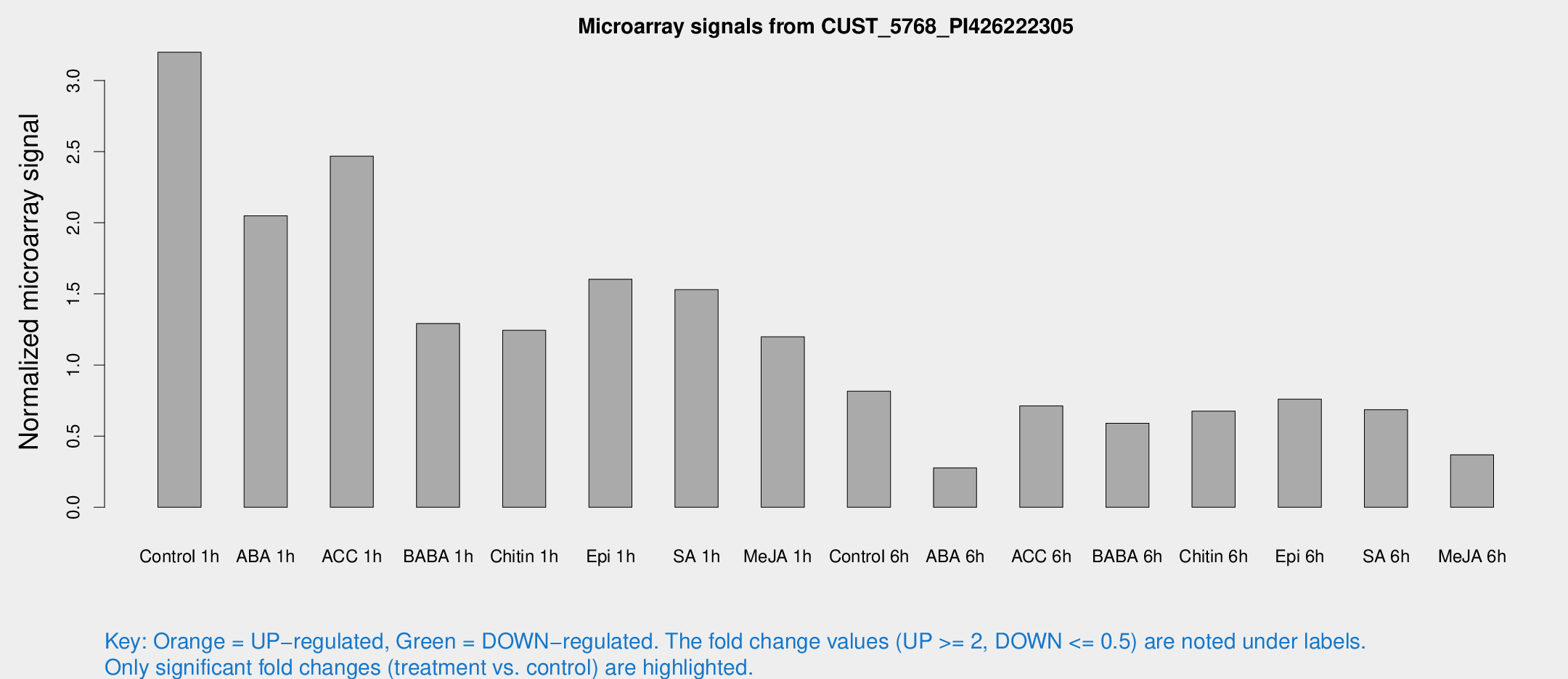
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 288.498 | 85.5148 | 3.19839 | 0.708927 |
| ABA 1h | 151.773 | 10.8028 | 2.0489 | 0.151124 |
| ACC 1h | 217.75 | 37.9984 | 2.46857 | 0.308348 |
| BABA 1h | 112.779 | 28.9556 | 1.29206 | 0.268208 |
| Chitin 1h | 96.0584 | 17.2942 | 1.24444 | 0.141115 |
| Epi 1h | 120.468 | 24.1404 | 1.60312 | 0.29632 |
| SA 1h | 148.568 | 47.5193 | 1.52966 | 0.613308 |
| Me-JA 1h | 81.9363 | 5.99714 | 1.1989 | 0.113127 |
| Control 6h | 71.9771 | 15.146 | 0.816291 | 0.126038 |
| ABA 6h | 25.475 | 4.83811 | 0.27663 | 0.050801 |
| ACC 6h | 72.0011 | 15.2564 | 0.712767 | 0.149048 |
| BABA 6h | 55.3205 | 5.33051 | 0.59099 | 0.0573195 |
| Chitin 6h | 64.5956 | 18.3106 | 0.675946 | 0.16665 |
| Epi 6h | 72.1819 | 7.72375 | 0.759671 | 0.0764672 |
| SA 6h | 59.3333 | 12.2699 | 0.685933 | 0.0824028 |
| Me-JA 6h | 32.1383 | 7.38031 | 0.36882 | 0.0534632 |
Source Transcript PGSC0003DMT400023124 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G52720.1 | +1 | 6e-77 | 242 | 120/239 (50%) | alpha carbonic anhydrase 1 | chr3:19538804-19541116 REVERSE LENGTH=284 |
