Probe CUST_5481_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5481_PI426222305 | JHI_St_60k_v1 | DMT400023125 | CAACCCACCCCTTAATTAAATGGAAGGCAAATAAATGTGTACAAATATTCCCCCATAATA |
All Microarray Probes Designed to Gene DMG400008954
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5481_PI426222305 | JHI_St_60k_v1 | DMT400023125 | CAACCCACCCCTTAATTAAATGGAAGGCAAATAAATGTGTACAAATATTCCCCCATAATA |
| CUST_5768_PI426222305 | JHI_St_60k_v1 | DMT400023124 | CCTTTGAGTTATAGTAATGAAAATTGTGCAAGCTTGGCTTAAAGGTTTTGAACAGACGAA |
Microarray Signals from CUST_5481_PI426222305
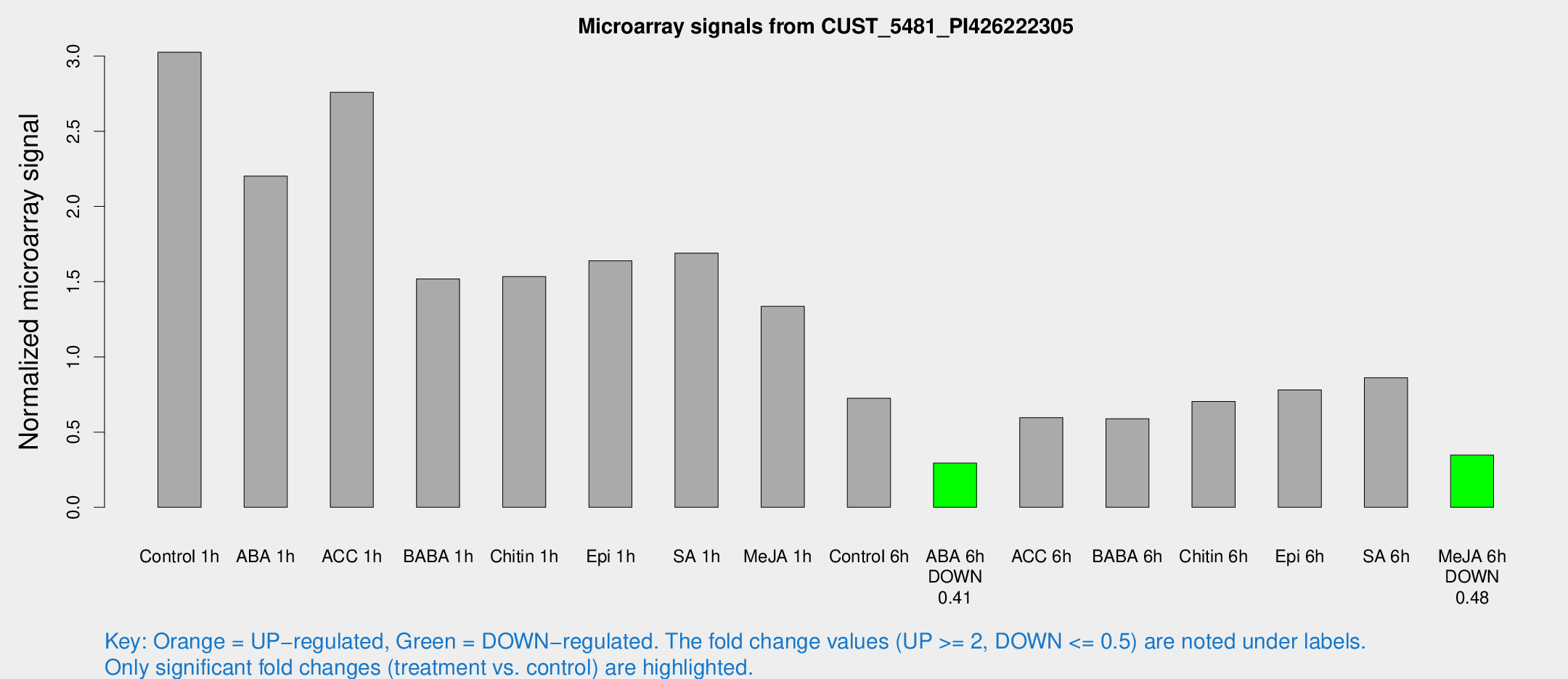
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2349.54 | 355.516 | 3.02523 | 0.372641 |
| ABA 1h | 1511.03 | 219.635 | 2.20127 | 0.42491 |
| ACC 1h | 2151.72 | 124.316 | 2.75831 | 0.251272 |
| BABA 1h | 1138.16 | 167.233 | 1.51815 | 0.0877759 |
| Chitin 1h | 1048.15 | 60.6148 | 1.53451 | 0.145892 |
| Epi 1h | 1116.94 | 195.902 | 1.63902 | 0.30772 |
| SA 1h | 1368.63 | 247.09 | 1.68906 | 0.357621 |
| Me-JA 1h | 835.161 | 77.7643 | 1.33568 | 0.20587 |
| Control 6h | 589.096 | 139.979 | 0.724894 | 0.136152 |
| ABA 6h | 237.759 | 14.1269 | 0.294157 | 0.0174624 |
| ACC 6h | 560.304 | 148.394 | 0.595781 | 0.10449 |
| BABA 6h | 512.696 | 79.6502 | 0.589133 | 0.0989511 |
| Chitin 6h | 572.076 | 43.9937 | 0.703732 | 0.0408584 |
| Epi 6h | 670.561 | 39.0139 | 0.780461 | 0.087397 |
| SA 6h | 649.967 | 49.0981 | 0.860602 | 0.0823915 |
| Me-JA 6h | 278.558 | 64.2033 | 0.347023 | 0.0588259 |
Source Transcript PGSC0003DMT400023125 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G52720.1 | +1 | 4e-77 | 242 | 120/239 (50%) | alpha carbonic anhydrase 1 | chr3:19538804-19541116 REVERSE LENGTH=284 |
