Probe CUST_5372_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5372_PI426222305 | JHI_St_60k_v1 | DMT400009327 | GTGTGACATATGTGAAAGTTCACCTGTTGATGGAAGTTCTCTTTGTCTGCAATGTGACAT |
All Microarray Probes Designed to Gene DMG400003625
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5372_PI426222305 | JHI_St_60k_v1 | DMT400009327 | GTGTGACATATGTGAAAGTTCACCTGTTGATGGAAGTTCTCTTTGTCTGCAATGTGACAT |
| CUST_5232_PI426222305 | JHI_St_60k_v1 | DMT400009328 | CCTTATTGGATCTCCATGCCTGGGAGAATTGTGAGTAGTGTGTTAAGTGTCGTTTATCAG |
Microarray Signals from CUST_5372_PI426222305
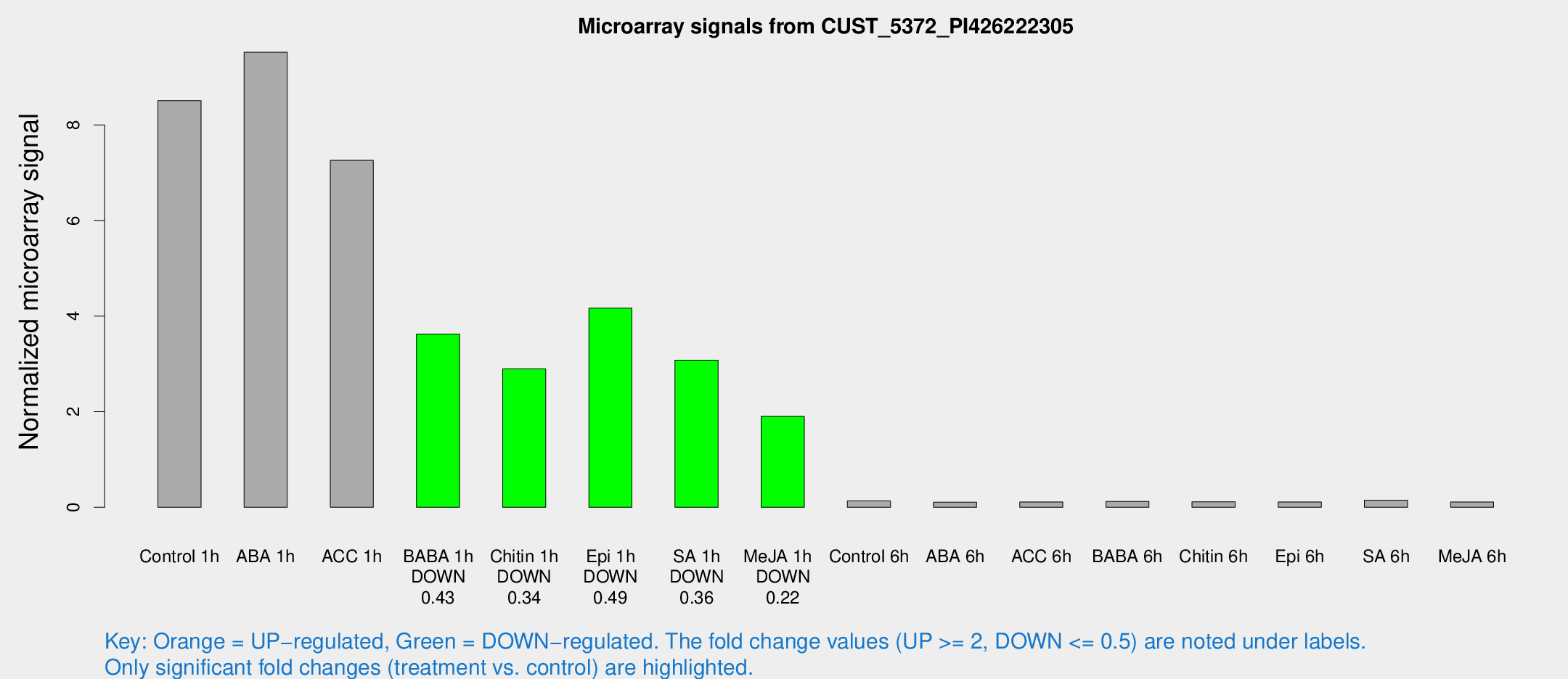
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 422.966 | 85.3896 | 8.50791 | 1.40932 |
| ABA 1h | 405.026 | 46.8777 | 9.52209 | 0.962625 |
| ACC 1h | 356.755 | 34.472 | 7.26013 | 0.593064 |
| BABA 1h | 179.523 | 44.3787 | 3.62516 | 0.723371 |
| Chitin 1h | 129.044 | 25.3957 | 2.89719 | 0.505941 |
| Epi 1h | 171.101 | 10.3156 | 4.16643 | 0.251168 |
| SA 1h | 153.801 | 24.8481 | 3.07996 | 0.300752 |
| Me-JA 1h | 73.7188 | 5.21755 | 1.90252 | 0.134467 |
| Control 6h | 6.41359 | 3.05721 | 0.13387 | 0.0649596 |
| ABA 6h | 5.45009 | 3.16409 | 0.10797 | 0.0626102 |
| ACC 6h | 6.21047 | 3.68698 | 0.111481 | 0.0645552 |
| BABA 6h | 6.55128 | 3.40462 | 0.122693 | 0.0641972 |
| Chitin 6h | 5.78609 | 3.35447 | 0.114567 | 0.0663995 |
| Epi 6h | 6.00755 | 3.51147 | 0.111411 | 0.0645757 |
| SA 6h | 7.39756 | 3.22204 | 0.148005 | 0.07251 |
| Me-JA 6h | 5.19089 | 3.00741 | 0.110042 | 0.0636179 |
Source Transcript PGSC0003DMT400009327 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G21320.1 | +1 | 2e-58 | 184 | 96/181 (53%) | B-box zinc finger family protein | chr2:9126502-9127652 FORWARD LENGTH=172 |
