Probe CUST_5232_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5232_PI426222305 | JHI_St_60k_v1 | DMT400009328 | CCTTATTGGATCTCCATGCCTGGGAGAATTGTGAGTAGTGTGTTAAGTGTCGTTTATCAG |
All Microarray Probes Designed to Gene DMG400003625
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5372_PI426222305 | JHI_St_60k_v1 | DMT400009327 | GTGTGACATATGTGAAAGTTCACCTGTTGATGGAAGTTCTCTTTGTCTGCAATGTGACAT |
| CUST_5232_PI426222305 | JHI_St_60k_v1 | DMT400009328 | CCTTATTGGATCTCCATGCCTGGGAGAATTGTGAGTAGTGTGTTAAGTGTCGTTTATCAG |
Microarray Signals from CUST_5232_PI426222305
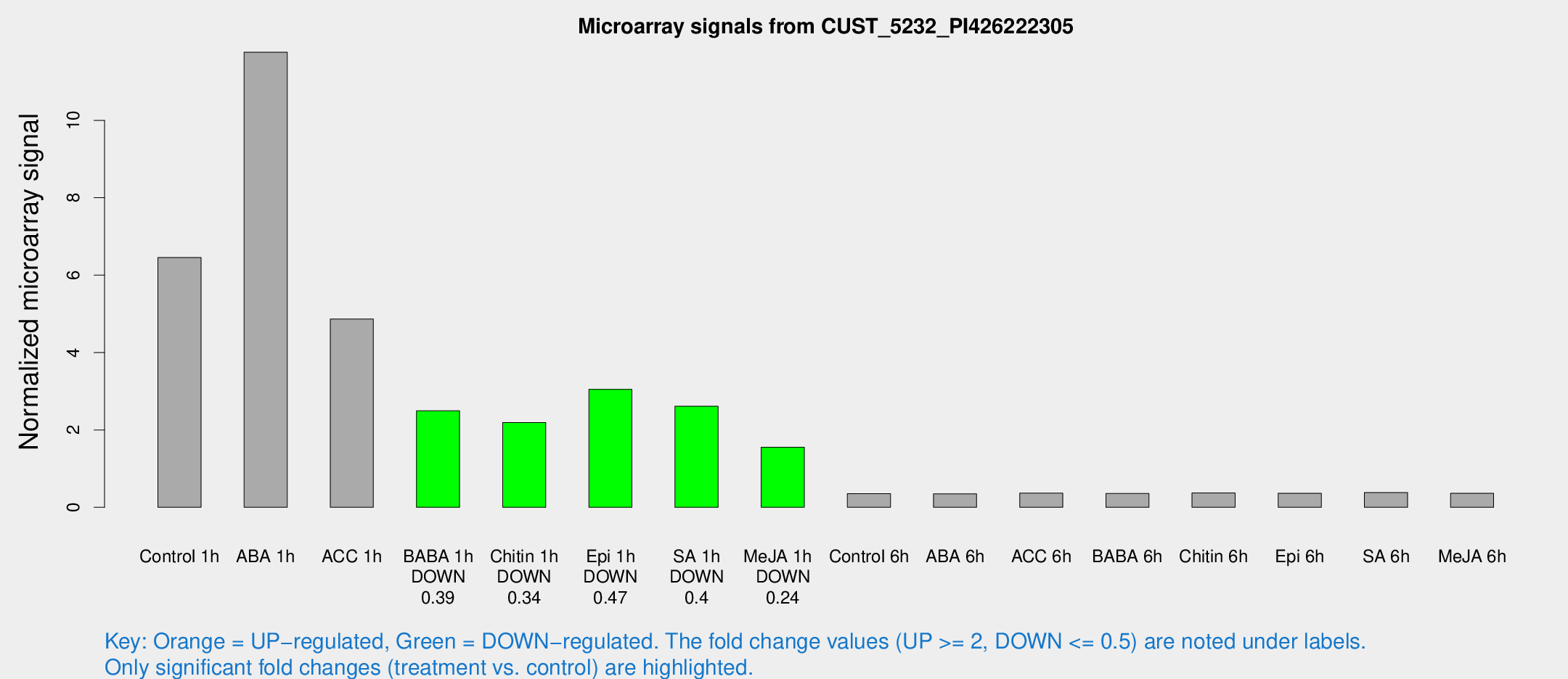
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 98.1153 | 20.4114 | 6.45368 | 0.899492 |
| ABA 1h | 153.354 | 16.1319 | 11.7602 | 2.12285 |
| ACC 1h | 73.42 | 6.65472 | 4.86535 | 0.498108 |
| BABA 1h | 36.1148 | 6.05526 | 2.49293 | 0.270348 |
| Chitin 1h | 30.7812 | 8.1992 | 2.19166 | 0.555927 |
| Epi 1h | 38.5022 | 3.64586 | 3.04841 | 0.288308 |
| SA 1h | 39.2868 | 3.71619 | 2.60977 | 0.248257 |
| Me-JA 1h | 18.561 | 3.18589 | 1.55315 | 0.266192 |
| Control 6h | 5.18973 | 3.00623 | 0.355178 | 0.205718 |
| ABA 6h | 5.39133 | 3.12488 | 0.347547 | 0.201251 |
| ACC 6h | 6.236 | 3.69268 | 0.365127 | 0.211468 |
| BABA 6h | 5.82637 | 3.38115 | 0.356323 | 0.20634 |
| Chitin 6h | 5.76045 | 3.337 | 0.371112 | 0.214912 |
| Epi 6h | 5.99443 | 3.49745 | 0.362024 | 0.209632 |
| SA 6h | 5.49429 | 3.18546 | 0.380823 | 0.22065 |
| Me-JA 6h | 5.20546 | 3.01691 | 0.36073 | 0.207476 |
Source Transcript PGSC0003DMT400009328 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G21320.1 | +2 | 3e-61 | 204 | 102/181 (56%) | B-box zinc finger family protein | chr2:9126502-9127652 FORWARD LENGTH=172 |
