Probe CUST_52788_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52788_PI426222305 | JHI_St_60k_v1 | DMT400081161 | TTGGCTGCAAACATTAAATTATTTCAGATGCTCACAGAGGAGTCAAATATGTTGGTCCAG |
All Microarray Probes Designed to Gene DMG400031717
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52788_PI426222305 | JHI_St_60k_v1 | DMT400081161 | TTGGCTGCAAACATTAAATTATTTCAGATGCTCACAGAGGAGTCAAATATGTTGGTCCAG |
| CUST_52787_PI426222305 | JHI_St_60k_v1 | DMT400081162 | TTGGCTGCAAACATTAAATTATTTCAGATGCTCACAGAGGAGTCAAATATGTTGGTCCAG |
Microarray Signals from CUST_52788_PI426222305
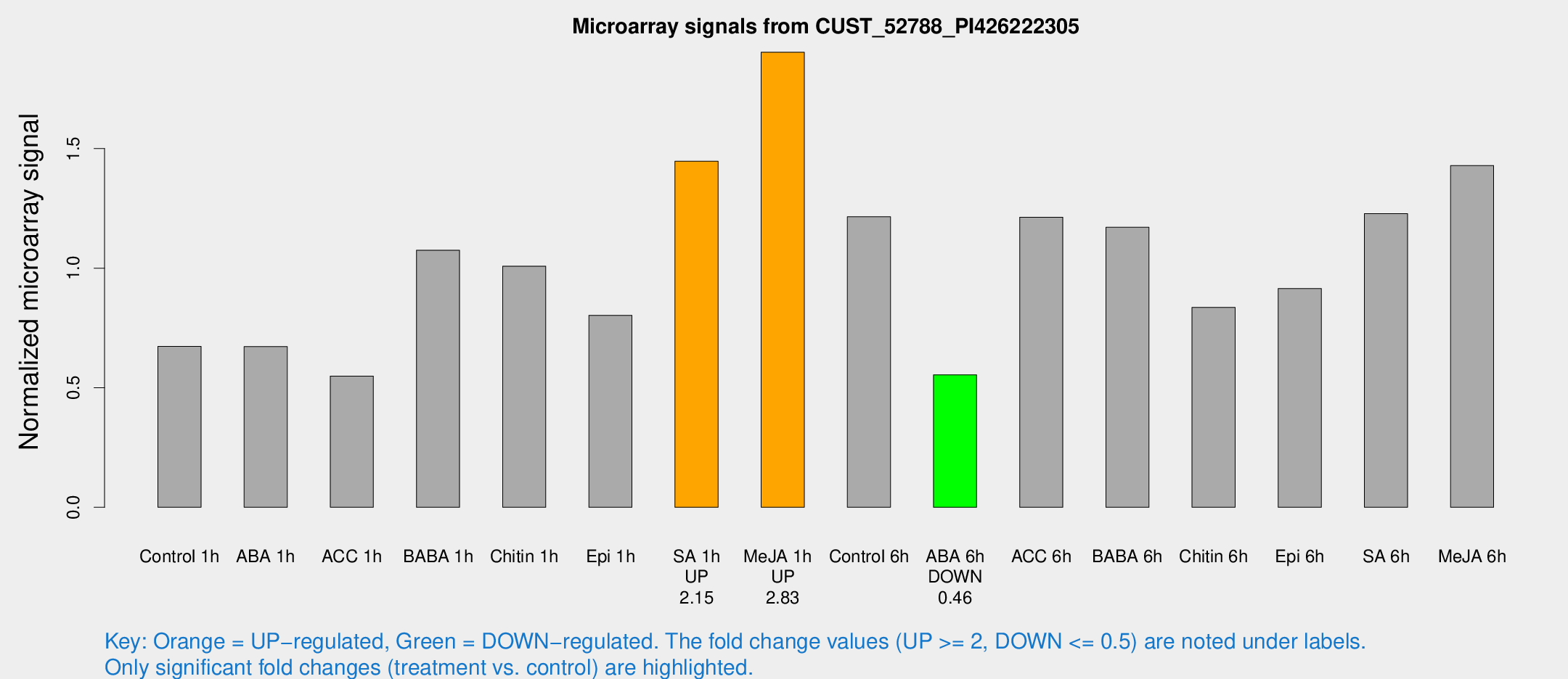
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 56.0663 | 8.30431 | 0.673106 | 0.101535 |
| ABA 1h | 50.6549 | 9.97744 | 0.671882 | 0.157426 |
| ACC 1h | 51.4776 | 18.2264 | 0.548864 | 0.193137 |
| BABA 1h | 89.1309 | 20.2125 | 1.07526 | 0.161835 |
| Chitin 1h | 73.9002 | 5.1985 | 1.00852 | 0.0707608 |
| Epi 1h | 59.5177 | 12.3322 | 0.802951 | 0.187136 |
| SA 1h | 125.568 | 25.0731 | 1.4474 | 0.318136 |
| Me-JA 1h | 127.372 | 10.8405 | 1.90307 | 0.119115 |
| Control 6h | 104.036 | 21.4219 | 1.21506 | 0.178819 |
| ABA 6h | 48.1416 | 4.2172 | 0.554103 | 0.0485983 |
| ACC 6h | 118.27 | 25.8579 | 1.21317 | 0.0928506 |
| BABA 6h | 107.112 | 7.19792 | 1.17104 | 0.0959252 |
| Chitin 6h | 75.3139 | 13.8921 | 0.836045 | 0.151235 |
| Epi 6h | 85.9756 | 13.0412 | 0.914876 | 0.0741993 |
| SA 6h | 103.71 | 21.0758 | 1.22817 | 0.142155 |
| Me-JA 6h | 116.511 | 9.10382 | 1.42919 | 0.0907731 |
Source Transcript PGSC0003DMT400081161 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G63120.1 | +1 | 1e-35 | 94 | 42/72 (58%) | cyclin p1;1 | chr3:23323143-23323893 REVERSE LENGTH=221 |
