Probe CUST_52787_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52787_PI426222305 | JHI_St_60k_v1 | DMT400081162 | TTGGCTGCAAACATTAAATTATTTCAGATGCTCACAGAGGAGTCAAATATGTTGGTCCAG |
All Microarray Probes Designed to Gene DMG400031717
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52788_PI426222305 | JHI_St_60k_v1 | DMT400081161 | TTGGCTGCAAACATTAAATTATTTCAGATGCTCACAGAGGAGTCAAATATGTTGGTCCAG |
| CUST_52787_PI426222305 | JHI_St_60k_v1 | DMT400081162 | TTGGCTGCAAACATTAAATTATTTCAGATGCTCACAGAGGAGTCAAATATGTTGGTCCAG |
Microarray Signals from CUST_52787_PI426222305
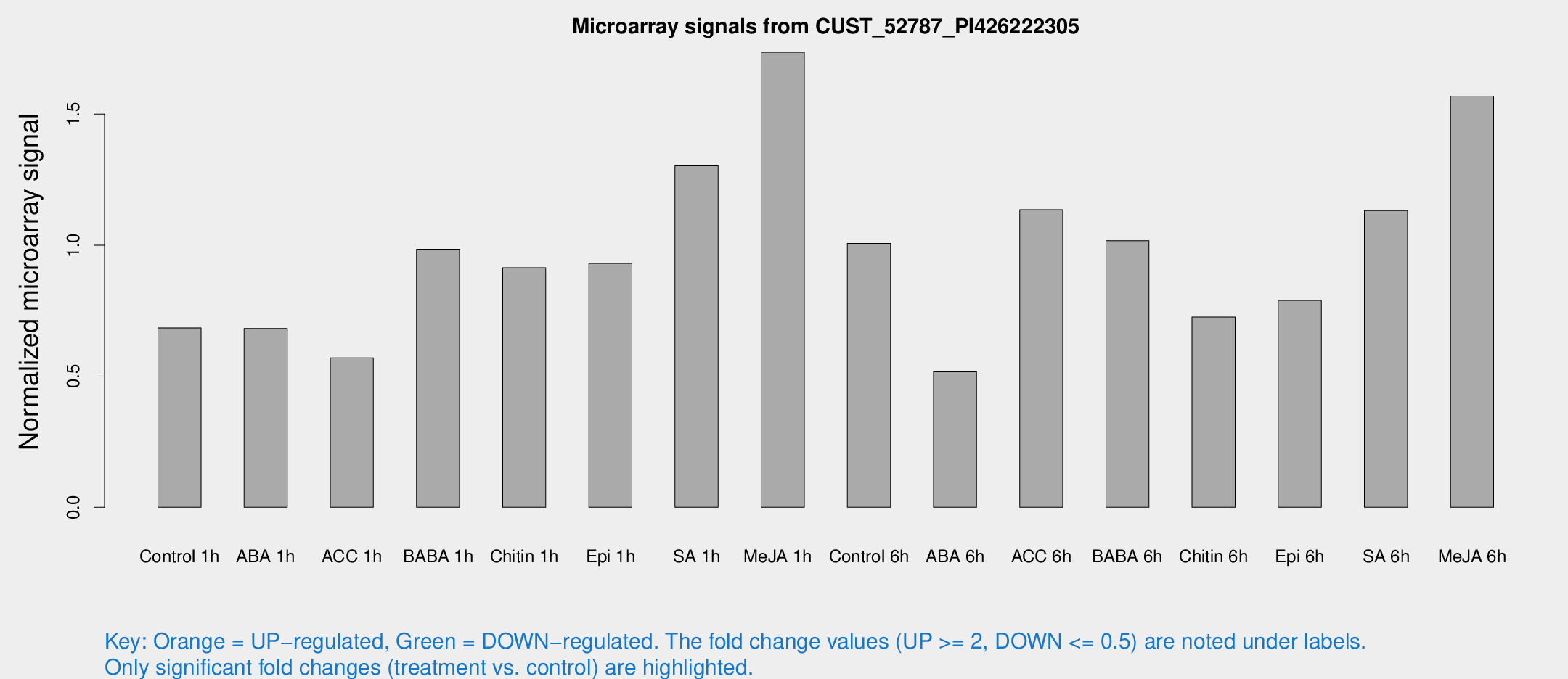
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 58.0041 | 8.01368 | 0.684412 | 0.0591549 |
| ABA 1h | 53.1367 | 12.9203 | 0.682356 | 0.132973 |
| ACC 1h | 50.9245 | 10.7045 | 0.570159 | 0.0915928 |
| BABA 1h | 81.1926 | 12.7037 | 0.984252 | 0.0761029 |
| Chitin 1h | 71.6672 | 14.0765 | 0.913861 | 0.149083 |
| Epi 1h | 69.0689 | 11.1051 | 0.930601 | 0.156514 |
| SA 1h | 113.987 | 17.7424 | 1.30277 | 0.190405 |
| Me-JA 1h | 126.353 | 29.4809 | 1.73579 | 0.360011 |
| Control 6h | 89.7043 | 20.7613 | 1.00671 | 0.185764 |
| ABA 6h | 45.9338 | 4.6019 | 0.516845 | 0.0611631 |
| ACC 6h | 109.398 | 10.5086 | 1.13505 | 0.0803212 |
| BABA 6h | 94.834 | 6.97924 | 1.01741 | 0.0747734 |
| Chitin 6h | 66.0272 | 11.135 | 0.725844 | 0.0978051 |
| Epi 6h | 74.8941 | 7.5686 | 0.789237 | 0.0645384 |
| SA 6h | 96.4827 | 17.0371 | 1.1319 | 0.0914391 |
| Me-JA 6h | 130.441 | 8.44262 | 1.5683 | 0.183871 |
Source Transcript PGSC0003DMT400081162 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G63120.1 | +1 | 5e-48 | 161 | 75/135 (56%) | cyclin p1;1 | chr3:23323143-23323893 REVERSE LENGTH=221 |
