Probe CUST_52186_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52186_PI426222305 | JHI_St_60k_v1 | DMT400080827 | TTGAAATCGGACGTGAAGTATCGTTTTGTGCTTGACATTGGCAACACATTGAACAAGAAT |
All Microarray Probes Designed to Gene DMG400031479
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52183_PI426222305 | JHI_St_60k_v1 | DMT400080830 | TTAGTAAGAAAGACGAAGCAATTGATCATTTAGGGGCAGACTCCTTTTGGTCAGTCGTGA |
| CUST_52186_PI426222305 | JHI_St_60k_v1 | DMT400080827 | TTGAAATCGGACGTGAAGTATCGTTTTGTGCTTGACATTGGCAACACATTGAACAAGAAT |
Microarray Signals from CUST_52186_PI426222305
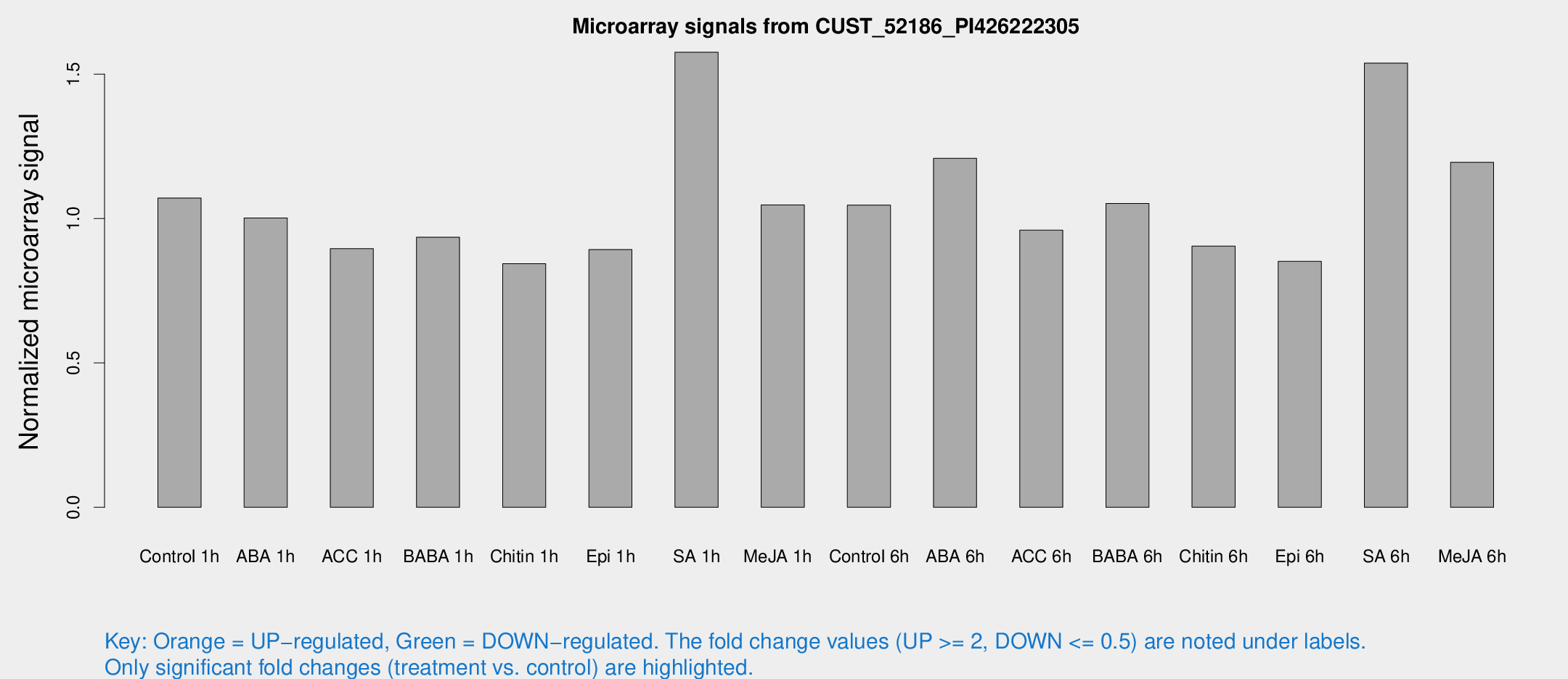
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4658.11 | 745.357 | 1.07113 | 0.101408 |
| ABA 1h | 3782.23 | 251.899 | 1.00232 | 0.0706652 |
| ACC 1h | 4057.78 | 739.079 | 0.895804 | 0.116189 |
| BABA 1h | 3928.57 | 597.993 | 0.935369 | 0.0618112 |
| Chitin 1h | 3254.72 | 338.51 | 0.843551 | 0.0487103 |
| Epi 1h | 3327.03 | 377.34 | 0.892574 | 0.0923005 |
| SA 1h | 6950.54 | 709.383 | 1.57623 | 0.129891 |
| Me-JA 1h | 3709.35 | 554.378 | 1.04702 | 0.0643875 |
| Control 6h | 4783.29 | 1137.52 | 1.0468 | 0.197657 |
| ABA 6h | 5519.62 | 597.907 | 1.20886 | 0.0697987 |
| ACC 6h | 4755.36 | 624.055 | 0.959735 | 0.0554167 |
| BABA 6h | 5007.87 | 289.5 | 1.05218 | 0.0607534 |
| Chitin 6h | 4097.89 | 237.053 | 0.904807 | 0.0522465 |
| Epi 6h | 4097.91 | 272.555 | 0.851999 | 0.0549272 |
| SA 6h | 6592.87 | 919.876 | 1.53847 | 0.10557 |
| Me-JA 6h | 5066.05 | 293.216 | 1.19499 | 0.102145 |
Source Transcript PGSC0003DMT400080827 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37990.1 | +1 | 1e-81 | 251 | 148/238 (62%) | elicitor-activated gene 3-2 | chr4:17855964-17857388 FORWARD LENGTH=359 |
