Probe CUST_52183_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52183_PI426222305 | JHI_St_60k_v1 | DMT400080830 | TTAGTAAGAAAGACGAAGCAATTGATCATTTAGGGGCAGACTCCTTTTGGTCAGTCGTGA |
All Microarray Probes Designed to Gene DMG400031479
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52183_PI426222305 | JHI_St_60k_v1 | DMT400080830 | TTAGTAAGAAAGACGAAGCAATTGATCATTTAGGGGCAGACTCCTTTTGGTCAGTCGTGA |
| CUST_52186_PI426222305 | JHI_St_60k_v1 | DMT400080827 | TTGAAATCGGACGTGAAGTATCGTTTTGTGCTTGACATTGGCAACACATTGAACAAGAAT |
Microarray Signals from CUST_52183_PI426222305
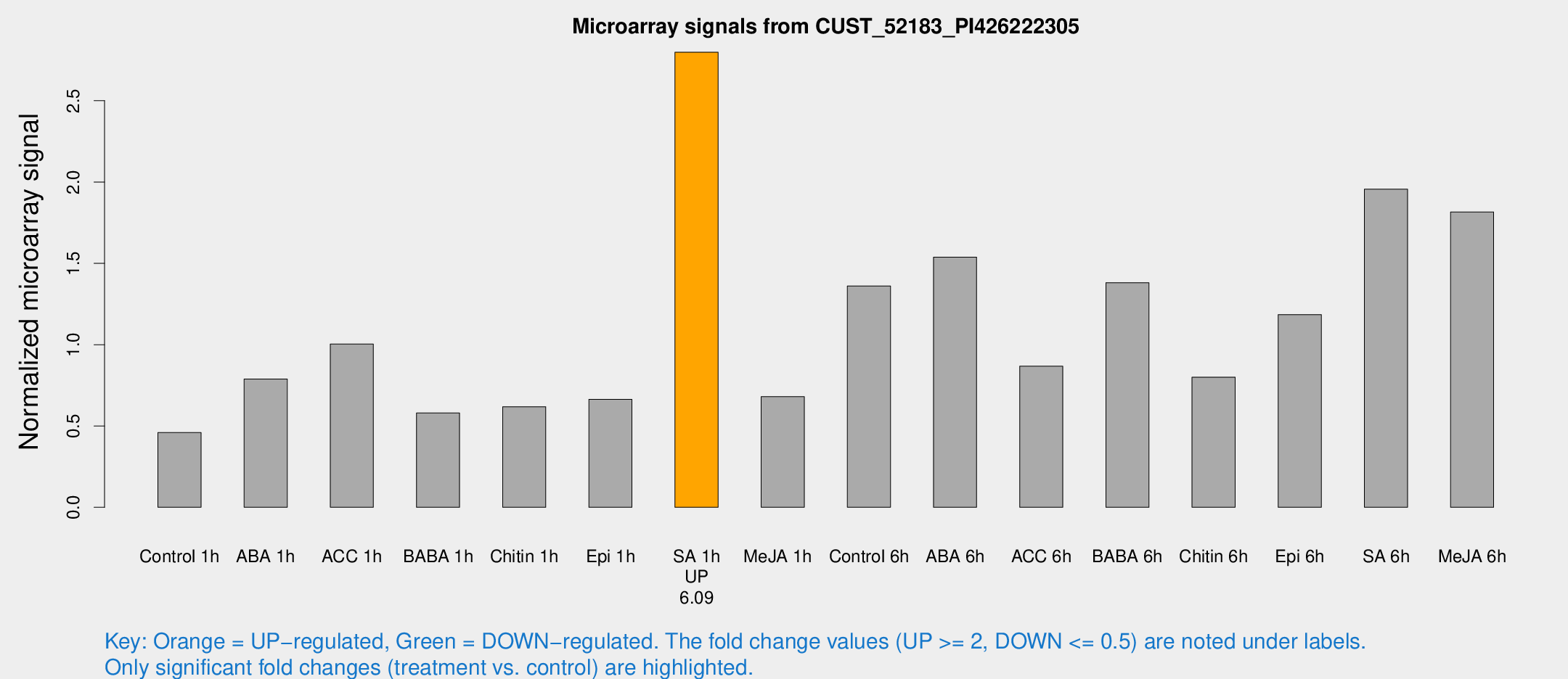
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13.9383 | 4.47794 | 0.459503 | 0.149737 |
| ABA 1h | 21.1535 | 6.78456 | 0.788926 | 0.260768 |
| ACC 1h | 29.1513 | 4.80187 | 1.0035 | 0.240309 |
| BABA 1h | 17.6609 | 6.31985 | 0.579856 | 0.177345 |
| Chitin 1h | 16.4151 | 4.36299 | 0.618569 | 0.158349 |
| Epi 1h | 20.1241 | 7.48426 | 0.663722 | 0.453371 |
| SA 1h | 79.4871 | 5.75407 | 2.7983 | 0.203234 |
| Me-JA 1h | 16.724 | 4.82538 | 0.680314 | 0.258909 |
| Control 6h | 47.412 | 22.5447 | 1.36116 | 0.610073 |
| ABA 6h | 48.0918 | 11.8349 | 1.53824 | 0.360959 |
| ACC 6h | 29.4852 | 8.07835 | 0.867463 | 0.148984 |
| BABA 6h | 50.7989 | 17.002 | 1.38116 | 0.778054 |
| Chitin 6h | 24.0885 | 4.5051 | 0.800489 | 0.15623 |
| Epi 6h | 37.8283 | 5.94829 | 1.1847 | 0.156986 |
| SA 6h | 87.4424 | 45.78 | 1.95611 | 2.08278 |
| Me-JA 6h | 50.8902 | 6.90648 | 1.81575 | 0.37524 |
Source Transcript PGSC0003DMT400080830 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37990.1 | +3 | 1e-42 | 99 | 65/87 (75%) | elicitor-activated gene 3-2 | chr4:17855964-17857388 FORWARD LENGTH=359 |
