Probe CUST_51143_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51143_PI426222305 | JHI_St_60k_v1 | DMT400008547 | CCAAGCTCAATCTATGCCTTGTTTTATTTGCAAAGATAAAAACTTACACGCACTTACATC |
All Microarray Probes Designed to Gene DMG400003305
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51143_PI426222305 | JHI_St_60k_v1 | DMT400008547 | CCAAGCTCAATCTATGCCTTGTTTTATTTGCAAAGATAAAAACTTACACGCACTTACATC |
| CUST_51156_PI426222305 | JHI_St_60k_v1 | DMT400008548 | CCAAGCTCAATCTATGCCTTGTTTTATTTGCAAAGATAAAAACTTACACGCACTTACATC |
Microarray Signals from CUST_51143_PI426222305
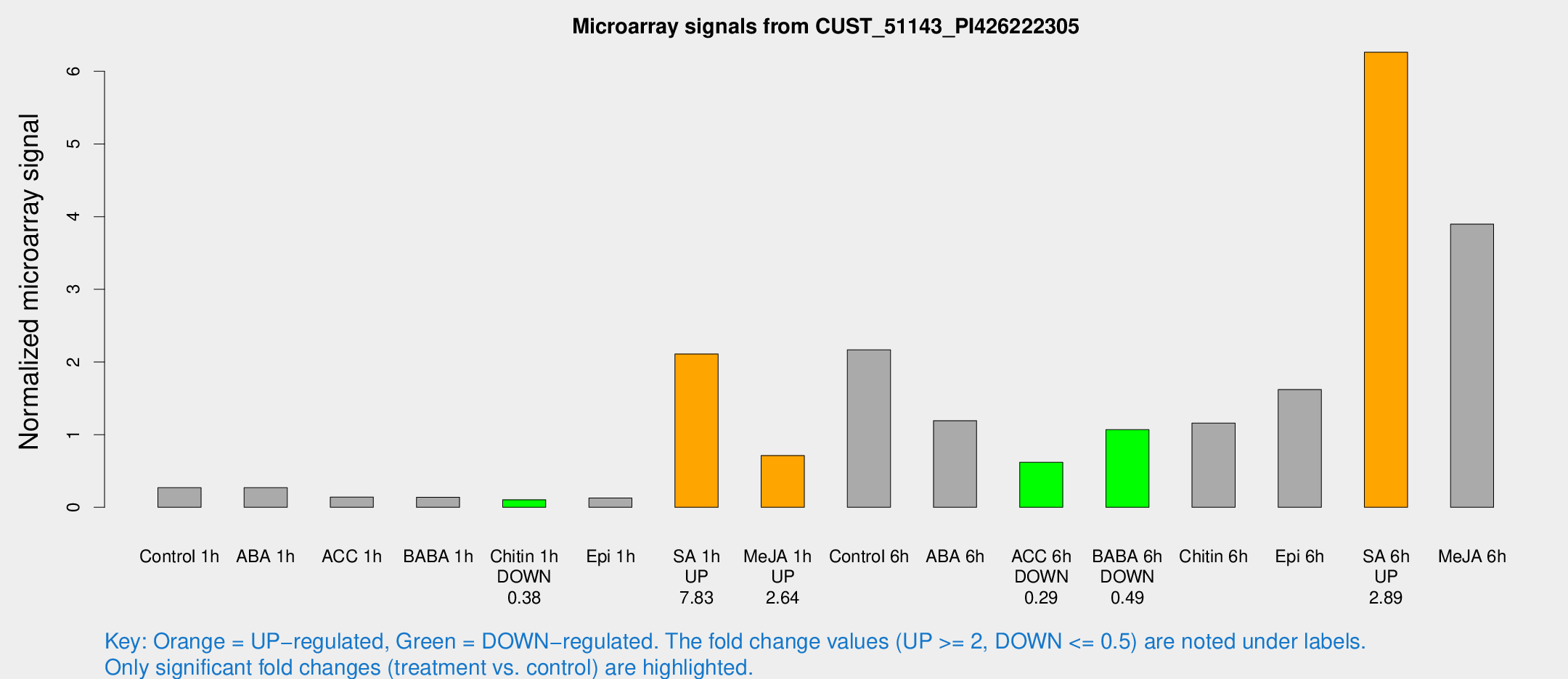
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 99.9615 | 22.4525 | 0.269607 | 0.0531372 |
| ABA 1h | 96.2288 | 29.11 | 0.270373 | 0.0987032 |
| ACC 1h | 79.9413 | 41.8315 | 0.141345 | 0.144327 |
| BABA 1h | 65.6809 | 27.2645 | 0.138495 | 0.101177 |
| Chitin 1h | 35.7221 | 10.2893 | 0.10376 | 0.0268482 |
| Epi 1h | 42.2262 | 11.3535 | 0.127165 | 0.0398061 |
| SA 1h | 781.42 | 108.365 | 2.1106 | 0.217977 |
| Me-JA 1h | 231.778 | 85.3424 | 0.712405 | 0.198624 |
| Control 6h | 774.735 | 80.1478 | 2.16816 | 0.125652 |
| ABA 6h | 452.955 | 53.1785 | 1.19249 | 0.079418 |
| ACC 6h | 254.922 | 33.0574 | 0.6197 | 0.160316 |
| BABA 6h | 424.241 | 33.5767 | 1.06818 | 0.083618 |
| Chitin 6h | 436.127 | 25.583 | 1.15929 | 0.0678785 |
| Epi 6h | 647.836 | 47.7092 | 1.62066 | 0.149622 |
| SA 6h | 2295.36 | 462.435 | 6.26262 | 0.70912 |
| Me-JA 6h | 1411.21 | 238.904 | 3.89771 | 0.695666 |
Source Transcript PGSC0003DMT400008547 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G10600.1 | +3 | 5e-126 | 384 | 215/477 (45%) | cytochrome P450, family 81, subfamily K, polypeptide 2 | chr5:3351227-3352777 FORWARD LENGTH=516 |
