Probe CUST_51156_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51156_PI426222305 | JHI_St_60k_v1 | DMT400008548 | CCAAGCTCAATCTATGCCTTGTTTTATTTGCAAAGATAAAAACTTACACGCACTTACATC |
All Microarray Probes Designed to Gene DMG400003305
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51143_PI426222305 | JHI_St_60k_v1 | DMT400008547 | CCAAGCTCAATCTATGCCTTGTTTTATTTGCAAAGATAAAAACTTACACGCACTTACATC |
| CUST_51156_PI426222305 | JHI_St_60k_v1 | DMT400008548 | CCAAGCTCAATCTATGCCTTGTTTTATTTGCAAAGATAAAAACTTACACGCACTTACATC |
Microarray Signals from CUST_51156_PI426222305
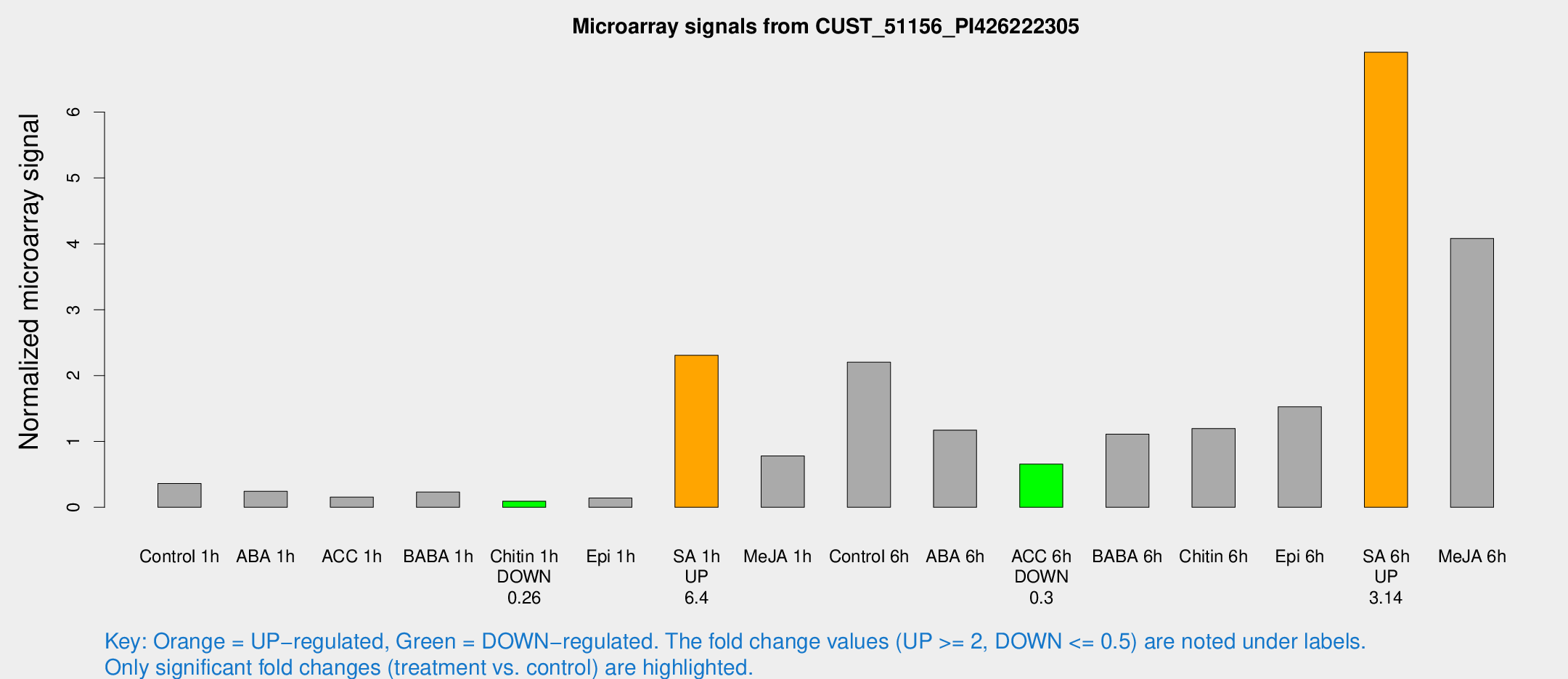
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 123.295 | 13.5698 | 0.360833 | 0.0233176 |
| ABA 1h | 83.3876 | 25.4359 | 0.243981 | 0.0927733 |
| ACC 1h | 81.2847 | 49.114 | 0.154685 | 0.126156 |
| BABA 1h | 86.783 | 27.0271 | 0.231841 | 0.0751747 |
| Chitin 1h | 29.5924 | 7.03402 | 0.0926596 | 0.0150662 |
| Epi 1h | 45.488 | 15.4051 | 0.14043 | 0.0484742 |
| SA 1h | 825.265 | 135.42 | 2.30779 | 0.308374 |
| Me-JA 1h | 235.239 | 74.5887 | 0.780123 | 0.177902 |
| Control 6h | 751.493 | 68.6045 | 2.20338 | 0.127738 |
| ABA 6h | 423.143 | 34.7379 | 1.17212 | 0.0685469 |
| ACC 6h | 257.114 | 28.9311 | 0.655597 | 0.168095 |
| BABA 6h | 425.727 | 52.0703 | 1.11121 | 0.118547 |
| Chitin 6h | 431.365 | 25.339 | 1.19713 | 0.0900174 |
| Epi 6h | 582.127 | 33.9909 | 1.52521 | 0.088776 |
| SA 6h | 2386.27 | 408.778 | 6.90949 | 0.497821 |
| Me-JA 6h | 1415.99 | 244.924 | 4.08285 | 0.740705 |
Source Transcript PGSC0003DMT400008548 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G10600.1 | +3 | 2e-118 | 366 | 217/514 (42%) | cytochrome P450, family 81, subfamily K, polypeptide 2 | chr5:3351227-3352777 FORWARD LENGTH=516 |
