Probe CUST_51091_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51091_PI426222305 | JHI_St_60k_v1 | DMT400077772 | TAGAGTTCAGCTGTTTCTTCGTGGATTCCTGTCTCATAAGCCGGAACTTCATAATGATAC |
All Microarray Probes Designed to Gene DMG400030253
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51090_PI426222305 | JHI_St_60k_v1 | DMT400077770 | TGGAATGGAAAGGTCGACAAAGTGAAGGGACAACTATATATTGCCTTCAACGCCAACCAT |
| CUST_51091_PI426222305 | JHI_St_60k_v1 | DMT400077772 | TAGAGTTCAGCTGTTTCTTCGTGGATTCCTGTCTCATAAGCCGGAACTTCATAATGATAC |
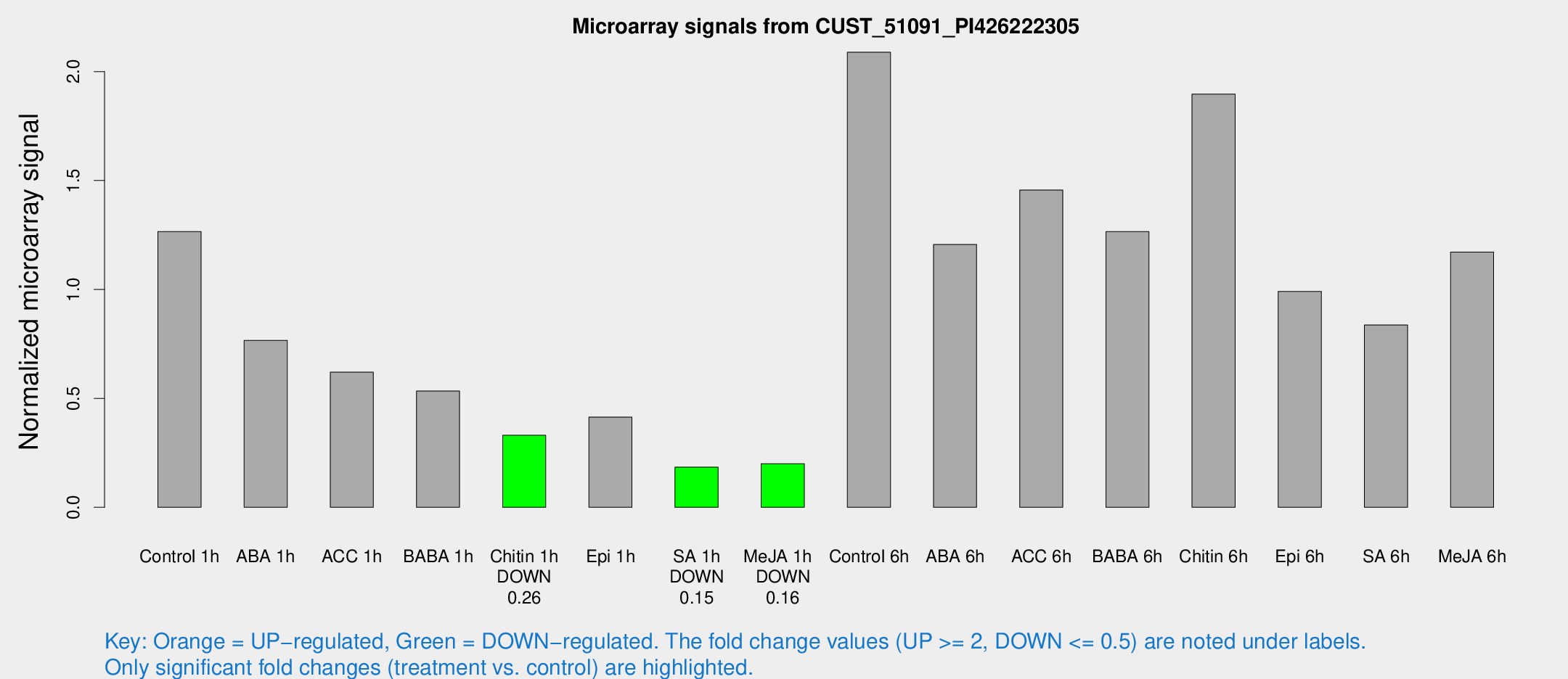
Microarray Signals from CUST_51091_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 44.6111 | 4.94474 | 1.266 | 0.118223 |
| ABA 1h | 25.9147 | 7.76423 | 0.766199 | 0.298594 |
| ACC 1h | 26.6831 | 9.27227 | 0.620063 | 0.277155 |
| BABA 1h | 19.8353 | 6.35418 | 0.533632 | 0.256474 |
| Chitin 1h | 11.5886 | 3.4284 | 0.330456 | 0.118693 |
| Epi 1h | 15.2289 | 6.53439 | 0.414168 | 0.220157 |
| SA 1h | 6.74746 | 3.24458 | 0.184267 | 0.0934211 |
| Me-JA 1h | 5.68799 | 3.30246 | 0.199685 | 0.11562 |
| Control 6h | 80.6872 | 22.0117 | 2.08889 | 0.542733 |
| ABA 6h | 45.5534 | 6.66714 | 1.20649 | 0.172037 |
| ACC 6h | 63.3657 | 18.9777 | 1.45637 | 0.512051 |
| BABA 6h | 54.4657 | 17.2753 | 1.26521 | 0.396227 |
| Chitin 6h | 71.8569 | 10.3026 | 1.89666 | 0.315242 |
| Epi 6h | 46.5121 | 16.0216 | 0.990654 | 0.646391 |
| SA 6h | 31.9489 | 9.79869 | 0.836762 | 0.424068 |
| Me-JA 6h | 46.3823 | 16.2027 | 1.17133 | 0.359924 |
Source Transcript PGSC0003DMT400077772 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G39930.1 | +1 | 7e-13 | 64 | 25/27 (93%) | isoamylase 1 | chr2:16666078-16672183 FORWARD LENGTH=783 |
