Probe CUST_51090_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51090_PI426222305 | JHI_St_60k_v1 | DMT400077770 | TGGAATGGAAAGGTCGACAAAGTGAAGGGACAACTATATATTGCCTTCAACGCCAACCAT |
All Microarray Probes Designed to Gene DMG400030253
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51090_PI426222305 | JHI_St_60k_v1 | DMT400077770 | TGGAATGGAAAGGTCGACAAAGTGAAGGGACAACTATATATTGCCTTCAACGCCAACCAT |
| CUST_51091_PI426222305 | JHI_St_60k_v1 | DMT400077772 | TAGAGTTCAGCTGTTTCTTCGTGGATTCCTGTCTCATAAGCCGGAACTTCATAATGATAC |
Microarray Signals from CUST_51090_PI426222305
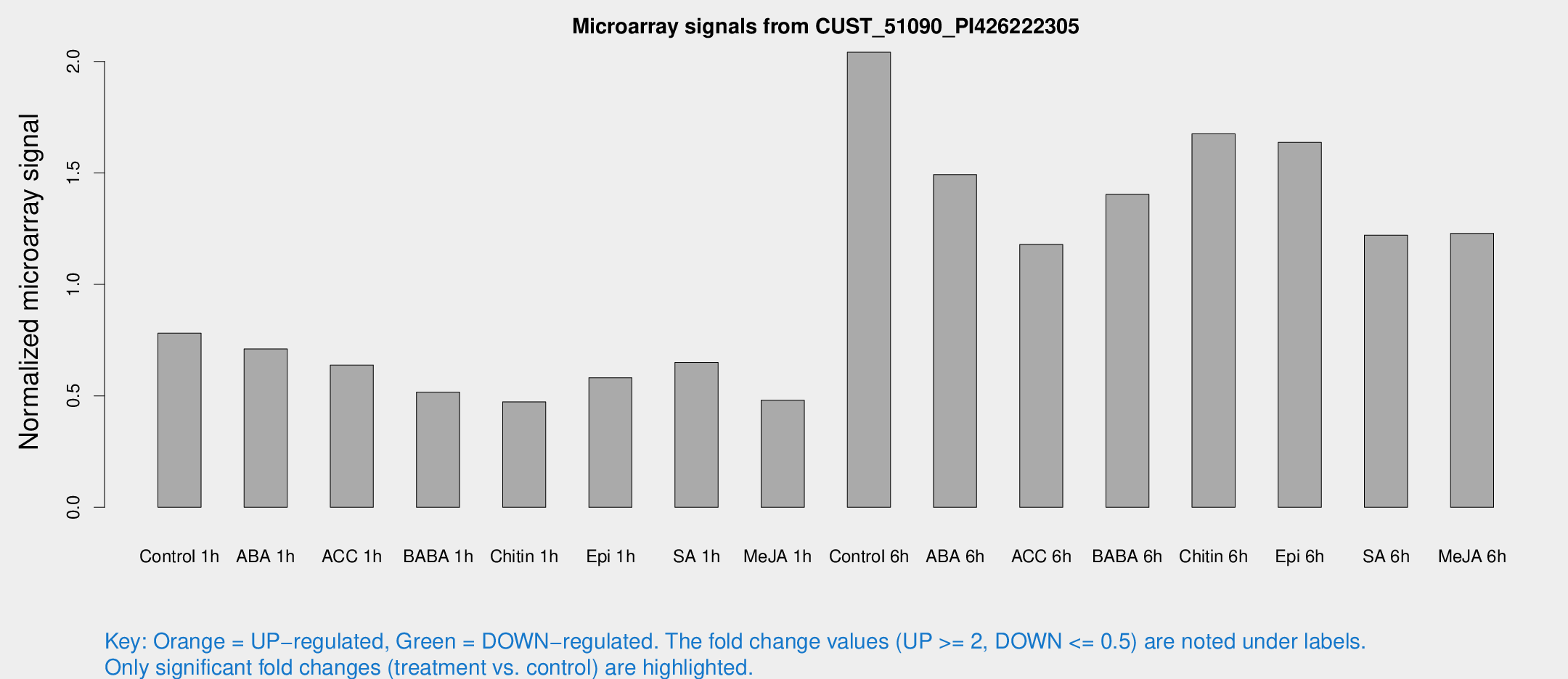
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 156.176 | 32.4228 | 0.781018 | 0.12641 |
| ABA 1h | 124.634 | 22.4307 | 0.710207 | 0.167652 |
| ACC 1h | 140.856 | 44.1545 | 0.637682 | 0.191438 |
| BABA 1h | 101.226 | 21.8696 | 0.516509 | 0.0762355 |
| Chitin 1h | 84.3725 | 15.8824 | 0.472979 | 0.0511621 |
| Epi 1h | 101.683 | 20.9676 | 0.580928 | 0.133747 |
| SA 1h | 129.642 | 12.8915 | 0.650352 | 0.0407361 |
| Me-JA 1h | 77.1317 | 11.5173 | 0.480257 | 0.037991 |
| Control 6h | 411.442 | 82.1581 | 2.04119 | 0.284385 |
| ABA 6h | 304.931 | 17.925 | 1.49186 | 0.0875817 |
| ACC 6h | 263.971 | 32.1336 | 1.17904 | 0.140092 |
| BABA 6h | 309.277 | 49.2509 | 1.40324 | 0.249226 |
| Chitin 6h | 344.389 | 27.0323 | 1.67479 | 0.098199 |
| Epi 6h | 361.887 | 49.4563 | 1.63719 | 0.147564 |
| SA 6h | 245.024 | 59.1742 | 1.22079 | 0.200051 |
| Me-JA 6h | 254.139 | 75.1494 | 1.22844 | 0.262287 |
Source Transcript PGSC0003DMT400077770 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G39930.1 | +1 | 3e-27 | 107 | 47/83 (57%) | isoamylase 1 | chr2:16666078-16672183 FORWARD LENGTH=783 |
