Probe CUST_48936_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48936_PI426222305 | JHI_St_60k_v1 | DMT400065309 | CGATCGAGTATGCAGTGGCGGATTTAGAATTTTAATTGATTTGAAGTTCGAGTTGTATAT |
All Microarray Probes Designed to Gene DMG400025389
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48933_PI426222305 | JHI_St_60k_v1 | DMT400065310 | CGATCGAGTATGCAGTGGCGGATTTAGAATTTTAATTGATTTGAAGTTCGAGTTGTATAT |
| CUST_48936_PI426222305 | JHI_St_60k_v1 | DMT400065309 | CGATCGAGTATGCAGTGGCGGATTTAGAATTTTAATTGATTTGAAGTTCGAGTTGTATAT |
Microarray Signals from CUST_48936_PI426222305
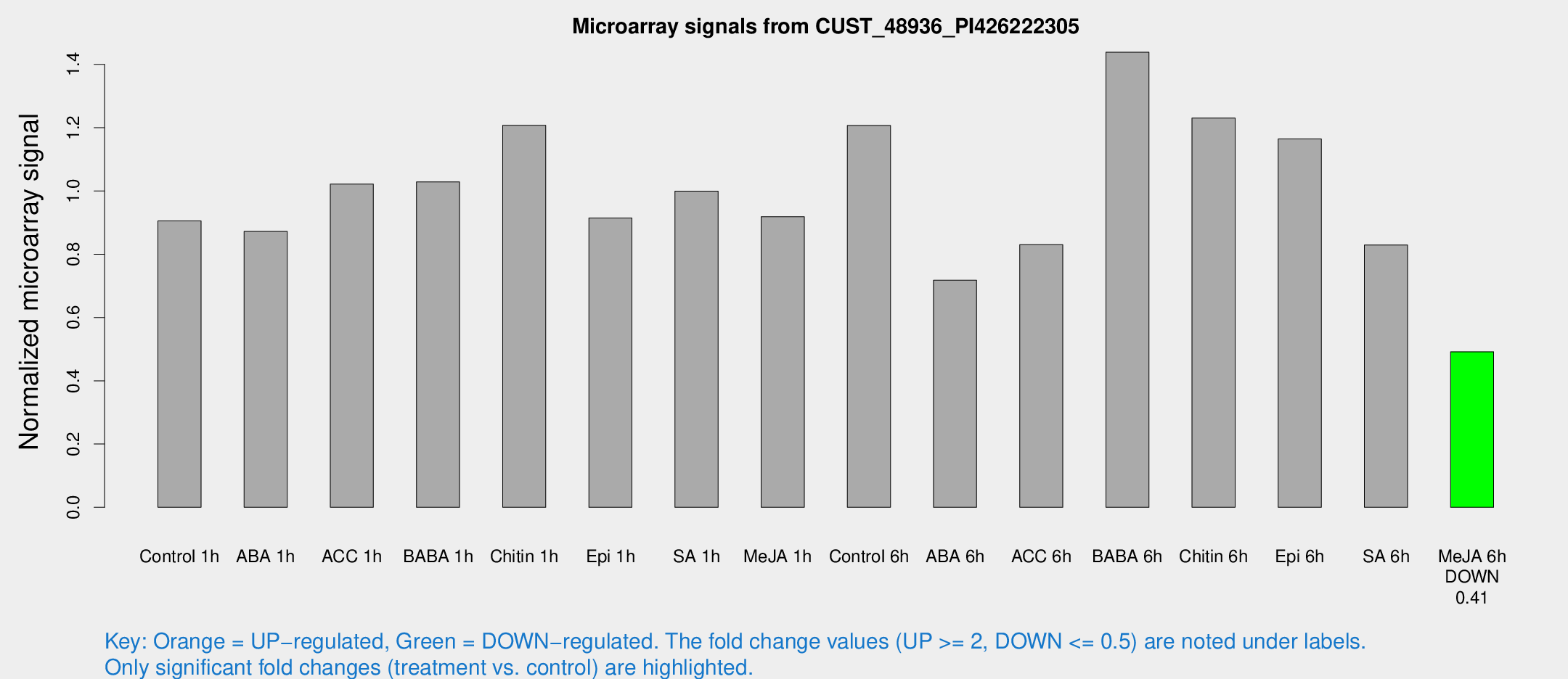
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 399.652 | 49.4055 | 0.905085 | 0.0778561 |
| ABA 1h | 342.932 | 52.1327 | 0.87192 | 0.119965 |
| ACC 1h | 514.544 | 161.344 | 1.02168 | 0.315487 |
| BABA 1h | 433.867 | 38.408 | 1.02852 | 0.0599549 |
| Chitin 1h | 483.439 | 78.3029 | 1.20731 | 0.183871 |
| Epi 1h | 352.237 | 55.2594 | 0.914615 | 0.127129 |
| SA 1h | 461.167 | 81.8104 | 0.999235 | 0.242736 |
| Me-JA 1h | 335.709 | 59.9468 | 0.918341 | 0.211615 |
| Control 6h | 565.873 | 137.769 | 1.20672 | 0.238234 |
| ABA 6h | 359.854 | 108.86 | 0.717635 | 0.252386 |
| ACC 6h | 425.304 | 73.3501 | 0.83026 | 0.0485637 |
| BABA 6h | 700.881 | 40.7347 | 1.43842 | 0.0963633 |
| Chitin 6h | 595.063 | 125.341 | 1.23014 | 0.256269 |
| Epi 6h | 572.382 | 36.1834 | 1.1644 | 0.156683 |
| SA 6h | 389.137 | 101.638 | 0.829192 | 0.1697 |
| Me-JA 6h | 217.237 | 32.3831 | 0.491468 | 0.0324277 |
Source Transcript PGSC0003DMT400065309 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G45650.1 | +1 | 2e-94 | 305 | 141/263 (54%) | nitrate excretion transporter1 | chr3:16759253-16761266 FORWARD LENGTH=558 |
