Probe CUST_48933_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48933_PI426222305 | JHI_St_60k_v1 | DMT400065310 | CGATCGAGTATGCAGTGGCGGATTTAGAATTTTAATTGATTTGAAGTTCGAGTTGTATAT |
All Microarray Probes Designed to Gene DMG400025389
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48933_PI426222305 | JHI_St_60k_v1 | DMT400065310 | CGATCGAGTATGCAGTGGCGGATTTAGAATTTTAATTGATTTGAAGTTCGAGTTGTATAT |
| CUST_48936_PI426222305 | JHI_St_60k_v1 | DMT400065309 | CGATCGAGTATGCAGTGGCGGATTTAGAATTTTAATTGATTTGAAGTTCGAGTTGTATAT |
Microarray Signals from CUST_48933_PI426222305
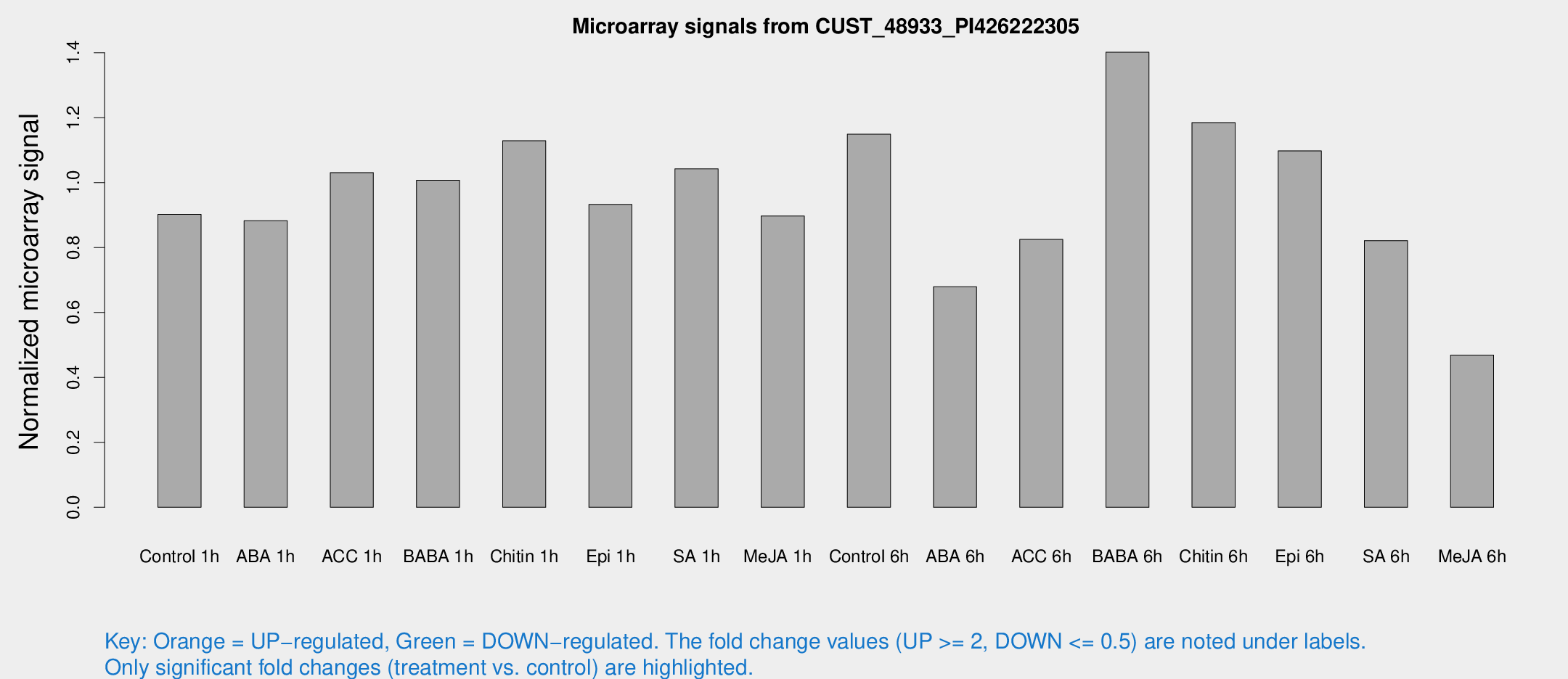
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 403.244 | 52.121 | 0.902357 | 0.0843423 |
| ABA 1h | 355.583 | 67.1437 | 0.882973 | 0.154878 |
| ACC 1h | 524.191 | 169.918 | 1.03097 | 0.312306 |
| BABA 1h | 429.997 | 43.4485 | 1.00722 | 0.111637 |
| Chitin 1h | 464.643 | 97.3781 | 1.12909 | 0.221521 |
| Epi 1h | 362.068 | 55.857 | 0.932807 | 0.113085 |
| SA 1h | 483.636 | 81.1728 | 1.04231 | 0.229888 |
| Me-JA 1h | 329.022 | 51.1968 | 0.897228 | 0.189916 |
| Control 6h | 557.105 | 151.141 | 1.14916 | 0.284431 |
| ABA 6h | 350.563 | 116.027 | 0.679318 | 0.2626 |
| ACC 6h | 424.437 | 65.4519 | 0.825 | 0.0482465 |
| BABA 6h | 692.895 | 56.1595 | 1.40171 | 0.118765 |
| Chitin 6h | 571.255 | 105.4 | 1.18481 | 0.20864 |
| Epi 6h | 549.035 | 55.2757 | 1.09803 | 0.199554 |
| SA 6h | 390.293 | 101.582 | 0.821604 | 0.170431 |
| Me-JA 6h | 214.155 | 45.1465 | 0.468903 | 0.0605273 |
Source Transcript PGSC0003DMT400065310 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G45650.1 | +1 | 0.0 | 529 | 286/542 (53%) | nitrate excretion transporter1 | chr3:16759253-16761266 FORWARD LENGTH=558 |
