Probe CUST_48904_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48904_PI426222305 | JHI_St_60k_v1 | DMT400056342 | ATCCTTACCCACCTTTGAACTAATATGAAGAGACAGATCGATGACTTATGTTGTTCGGAC |
All Microarray Probes Designed to Gene DMG400021890
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48915_PI426222305 | JHI_St_60k_v1 | DMT400056343 | ATCCTTACCCACCTTTGAACTAATATGAAGAGACAGATCGATGACTTATGTTGTTCGGAC |
| CUST_48904_PI426222305 | JHI_St_60k_v1 | DMT400056342 | ATCCTTACCCACCTTTGAACTAATATGAAGAGACAGATCGATGACTTATGTTGTTCGGAC |
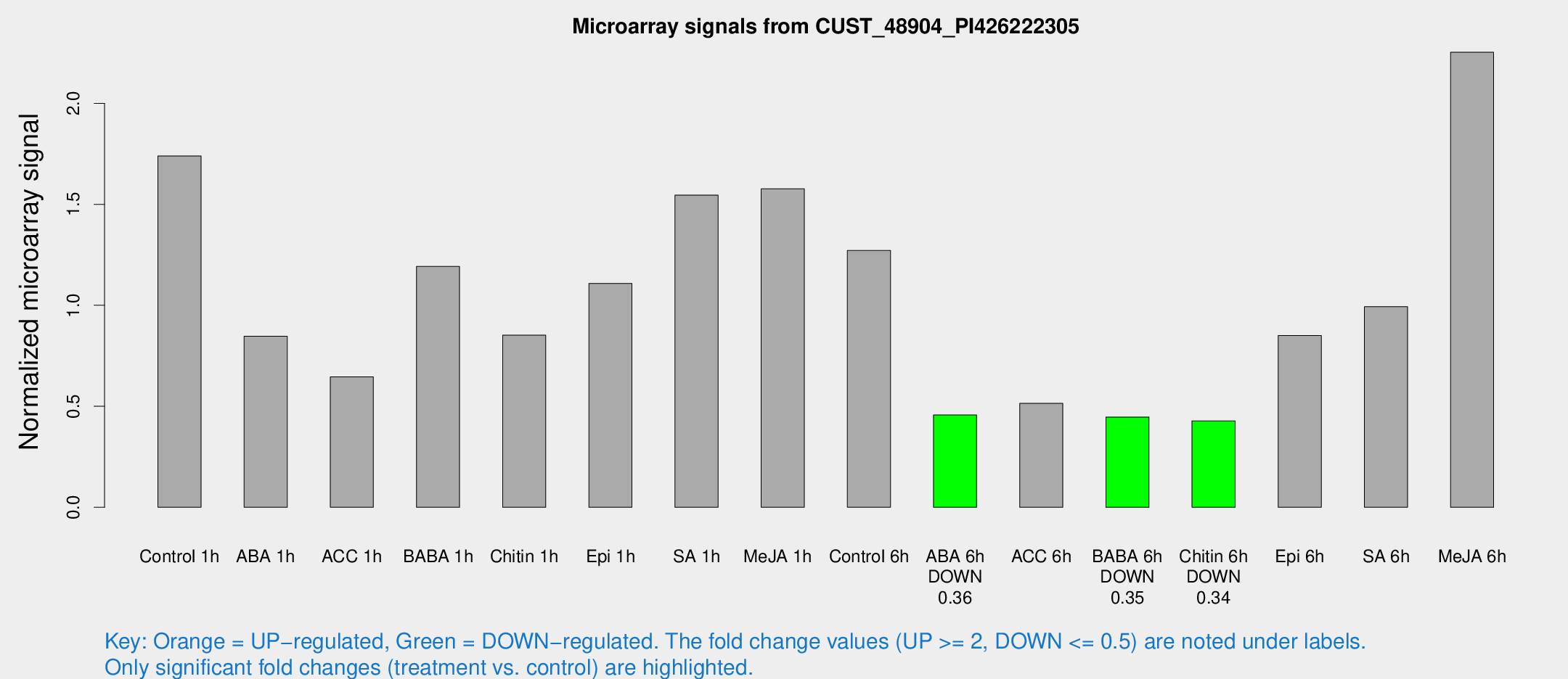
Microarray Signals from CUST_48904_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 496.397 | 99.0495 | 1.73896 | 0.440203 |
| ABA 1h | 208.909 | 24.4686 | 0.846859 | 0.0510917 |
| ACC 1h | 267.123 | 144.142 | 0.645856 | 0.516203 |
| BABA 1h | 328.417 | 62.6153 | 1.19215 | 0.132656 |
| Chitin 1h | 213.761 | 24.4492 | 0.852141 | 0.139811 |
| Epi 1h | 289.504 | 78.0087 | 1.10826 | 0.370749 |
| SA 1h | 487.371 | 169.766 | 1.54554 | 0.550982 |
| Me-JA 1h | 356.261 | 28.0034 | 1.57679 | 0.0924714 |
| Control 6h | 358.602 | 50.9357 | 1.27128 | 0.268937 |
| ABA 6h | 134.227 | 8.77194 | 0.457 | 0.0297947 |
| ACC 6h | 169.839 | 35.1709 | 0.514371 | 0.197099 |
| BABA 6h | 138.045 | 9.05841 | 0.446913 | 0.0292504 |
| Chitin 6h | 126.177 | 10.9073 | 0.4275 | 0.0521776 |
| Epi 6h | 272.778 | 47.6024 | 0.850864 | 0.209526 |
| SA 6h | 282.534 | 54.6245 | 0.993375 | 0.102168 |
| Me-JA 6h | 622.515 | 56.2788 | 2.2529 | 0.343552 |
Source Transcript PGSC0003DMT400056342 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G13700.1 | +3 | 7e-92 | 281 | 146/210 (70%) | 6-phosphogluconolactonase 1 | chr1:4694475-4695600 REVERSE LENGTH=268 |
