Probe CUST_48915_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48915_PI426222305 | JHI_St_60k_v1 | DMT400056343 | ATCCTTACCCACCTTTGAACTAATATGAAGAGACAGATCGATGACTTATGTTGTTCGGAC |
All Microarray Probes Designed to Gene DMG400021890
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48915_PI426222305 | JHI_St_60k_v1 | DMT400056343 | ATCCTTACCCACCTTTGAACTAATATGAAGAGACAGATCGATGACTTATGTTGTTCGGAC |
| CUST_48904_PI426222305 | JHI_St_60k_v1 | DMT400056342 | ATCCTTACCCACCTTTGAACTAATATGAAGAGACAGATCGATGACTTATGTTGTTCGGAC |
Microarray Signals from CUST_48915_PI426222305
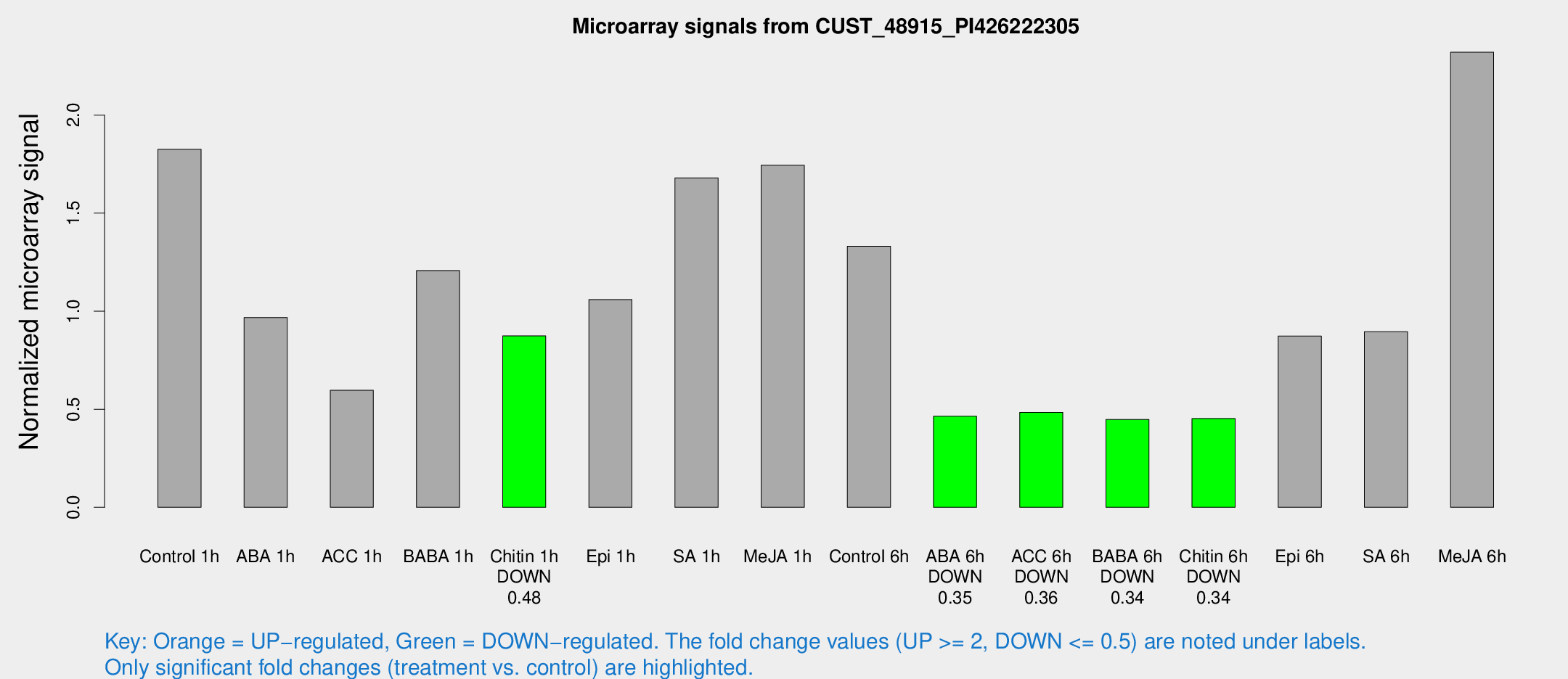
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 507.492 | 59.3 | 1.82556 | 0.315847 |
| ABA 1h | 237.077 | 22.0446 | 0.967795 | 0.057761 |
| ACC 1h | 254.32 | 134.977 | 0.59645 | 0.541129 |
| BABA 1h | 331.125 | 63.6948 | 1.20714 | 0.145828 |
| Chitin 1h | 219.59 | 27.8928 | 0.874059 | 0.146834 |
| Epi 1h | 264.434 | 55.3344 | 1.05954 | 0.242452 |
| SA 1h | 544.018 | 213.128 | 1.67964 | 0.680391 |
| Me-JA 1h | 392.466 | 24.9229 | 1.74472 | 0.102035 |
| Control 6h | 375.176 | 56.0163 | 1.33102 | 0.26913 |
| ABA 6h | 139.293 | 23.6122 | 0.464348 | 0.0514268 |
| ACC 6h | 155.342 | 20.8583 | 0.484363 | 0.13887 |
| BABA 6h | 137.662 | 8.98615 | 0.447552 | 0.0316061 |
| Chitin 6h | 133.733 | 13.6551 | 0.452745 | 0.0633248 |
| Epi 6h | 278.86 | 46.6781 | 0.873051 | 0.22044 |
| SA 6h | 247.044 | 30.4647 | 0.895206 | 0.0538369 |
| Me-JA 6h | 636.412 | 38.2518 | 2.32051 | 0.322451 |
Source Transcript PGSC0003DMT400056343 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G13700.1 | +3 | 5e-115 | 340 | 175/261 (67%) | 6-phosphogluconolactonase 1 | chr1:4694475-4695600 REVERSE LENGTH=268 |
