Probe CUST_48676_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48676_PI426222305 | JHI_St_60k_v1 | DMT400080319 | TGCTTTTTAGCACTGAGCAGTAGTTCCTGTGTTGGTAAATTTTGACTGATGAATCACAAA |
All Microarray Probes Designed to Gene DMG400031269
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48675_PI426222305 | JHI_St_60k_v1 | DMT400080317 | TGCTTTTTAGCACTGAGCAGTAGTTCCTGTGTTGGTAAATTTTGACTGATGAATCACAAA |
| CUST_48676_PI426222305 | JHI_St_60k_v1 | DMT400080319 | TGCTTTTTAGCACTGAGCAGTAGTTCCTGTGTTGGTAAATTTTGACTGATGAATCACAAA |
Microarray Signals from CUST_48676_PI426222305
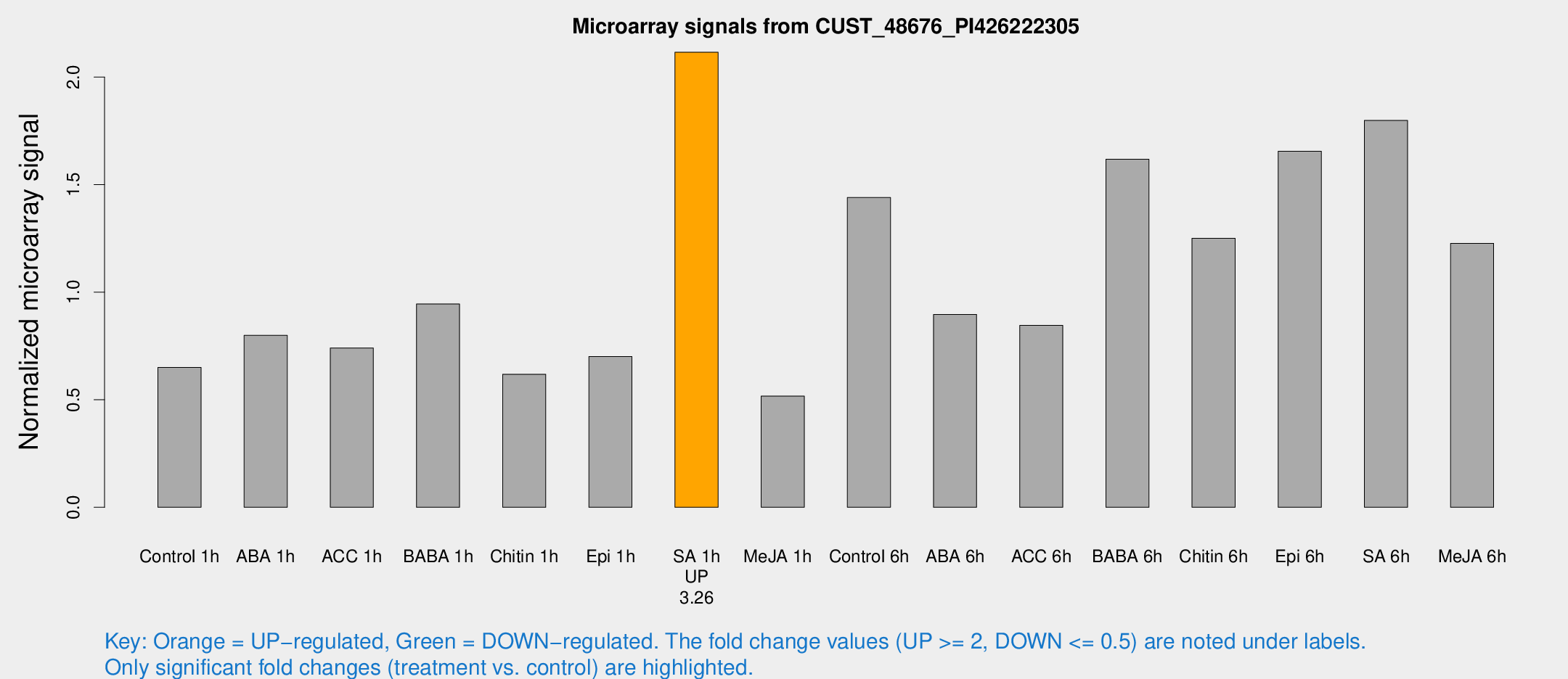
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 82.0346 | 10.4873 | 0.649853 | 0.0473743 |
| ABA 1h | 88.7596 | 9.49384 | 0.799567 | 0.0898921 |
| ACC 1h | 108.476 | 33.3384 | 0.740831 | 0.262385 |
| BABA 1h | 113.748 | 8.6576 | 0.945241 | 0.0629201 |
| Chitin 1h | 71.4567 | 12.4224 | 0.618518 | 0.0747672 |
| Epi 1h | 75.6193 | 5.595 | 0.701157 | 0.0522527 |
| SA 1h | 274.01 | 32.7358 | 2.11544 | 0.164159 |
| Me-JA 1h | 52.4506 | 4.80571 | 0.516667 | 0.0474918 |
| Control 6h | 190.437 | 45.8913 | 1.43987 | 0.238236 |
| ABA 6h | 118.889 | 8.50177 | 0.896519 | 0.0592172 |
| ACC 6h | 123.182 | 17.5579 | 0.845445 | 0.113387 |
| BABA 6h | 225.917 | 15.8043 | 1.61815 | 0.0979081 |
| Chitin 6h | 165.869 | 10.4188 | 1.25058 | 0.108302 |
| Epi 6h | 232.979 | 18.3147 | 1.65494 | 0.173267 |
| SA 6h | 225.702 | 31.6029 | 1.79846 | 0.108944 |
| Me-JA 6h | 166.778 | 54.1026 | 1.22694 | 0.29415 |
Source Transcript PGSC0003DMT400080319 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G14790.1 | +1 | 0.0 | 1290 | 680/1103 (62%) | RNA-dependent RNA polymerase 1 | chr1:5094317-5097817 REVERSE LENGTH=1107 |
