Probe CUST_48675_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48675_PI426222305 | JHI_St_60k_v1 | DMT400080317 | TGCTTTTTAGCACTGAGCAGTAGTTCCTGTGTTGGTAAATTTTGACTGATGAATCACAAA |
All Microarray Probes Designed to Gene DMG400031269
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48675_PI426222305 | JHI_St_60k_v1 | DMT400080317 | TGCTTTTTAGCACTGAGCAGTAGTTCCTGTGTTGGTAAATTTTGACTGATGAATCACAAA |
| CUST_48676_PI426222305 | JHI_St_60k_v1 | DMT400080319 | TGCTTTTTAGCACTGAGCAGTAGTTCCTGTGTTGGTAAATTTTGACTGATGAATCACAAA |
Microarray Signals from CUST_48675_PI426222305
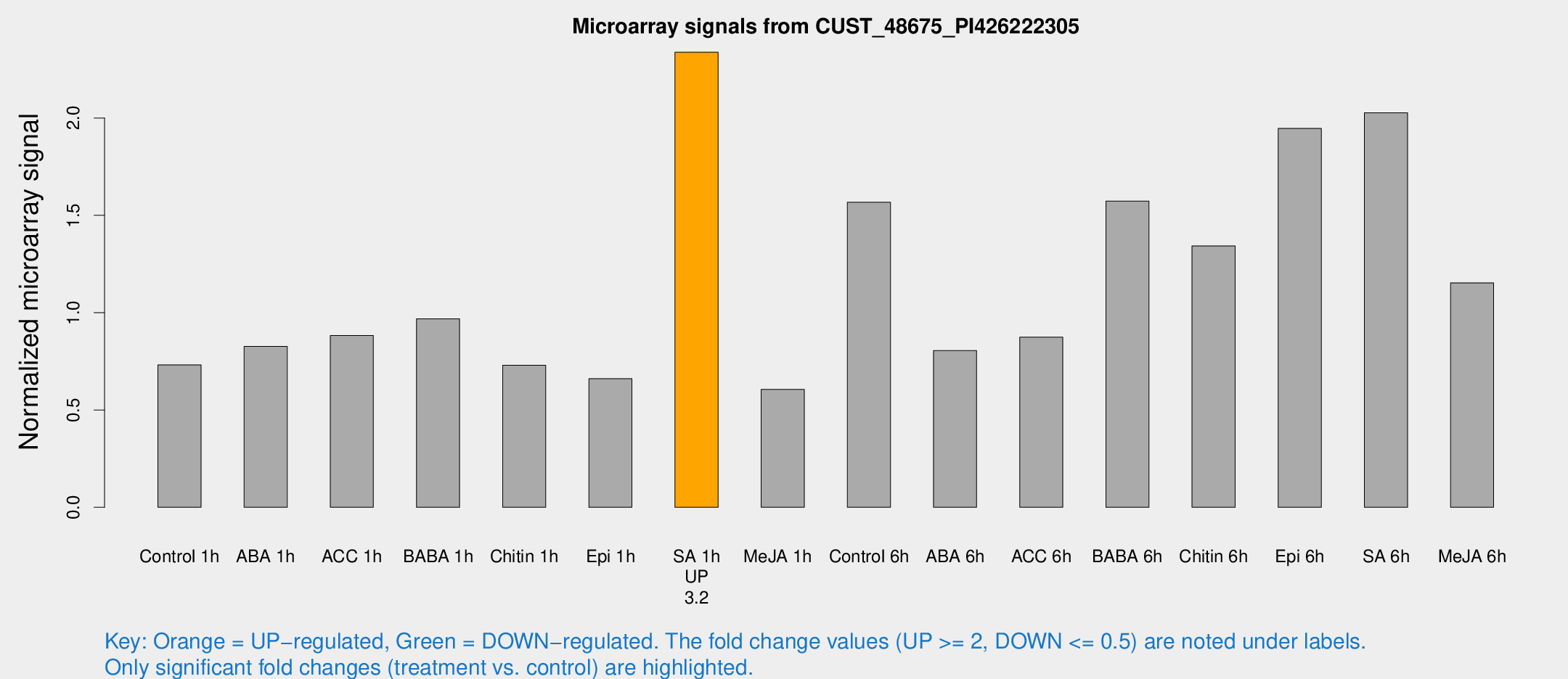
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 87.235 | 20.1413 | 0.731293 | 0.123516 |
| ABA 1h | 83.5343 | 6.21758 | 0.826398 | 0.0561517 |
| ACC 1h | 114.027 | 31.3039 | 0.882767 | 0.245741 |
| BABA 1h | 107.822 | 13.2608 | 0.968623 | 0.0638167 |
| Chitin 1h | 76.3243 | 11.112 | 0.730057 | 0.0848607 |
| Epi 1h | 65.7171 | 7.2611 | 0.660511 | 0.0830337 |
| SA 1h | 276.18 | 29.0794 | 2.33795 | 0.148896 |
| Me-JA 1h | 56.9306 | 6.23382 | 0.605896 | 0.048885 |
| Control 6h | 196.222 | 56.1999 | 1.56708 | 0.368146 |
| ABA 6h | 97.5936 | 6.53452 | 0.805091 | 0.0538494 |
| ACC 6h | 115.302 | 12.2162 | 0.874036 | 0.0799426 |
| BABA 6h | 201.165 | 14.7599 | 1.57331 | 0.0949876 |
| Chitin 6h | 163.741 | 15.0339 | 1.34265 | 0.166792 |
| Epi 6h | 250.96 | 20.5806 | 1.94637 | 0.231382 |
| SA 6h | 233.584 | 36.1483 | 2.02719 | 0.150272 |
| Me-JA 6h | 142.753 | 43.8734 | 1.15321 | 0.257633 |
Source Transcript PGSC0003DMT400080317 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G14790.1 | +2 | 7e-108 | 346 | 169/251 (67%) | RNA-dependent RNA polymerase 1 | chr1:5094317-5097817 REVERSE LENGTH=1107 |
