Probe CUST_48000_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48000_PI426222305 | JHI_St_60k_v1 | DMT400074454 | TTATCCTTTACAGGTTATTCGGACCAGAATGCAAGCTGATTCTGCATACAAAGGTATGGC |
All Microarray Probes Designed to Gene DMG402028933
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48003_PI426222305 | JHI_St_60k_v1 | DMT400074453 | TTTCTTCCCCTTCTATTTGGATGGTGTAGCTTTCTGGTTTATCATGTGACATGCCGTTTT |
| CUST_48000_PI426222305 | JHI_St_60k_v1 | DMT400074454 | TTATCCTTTACAGGTTATTCGGACCAGAATGCAAGCTGATTCTGCATACAAAGGTATGGC |
Microarray Signals from CUST_48000_PI426222305
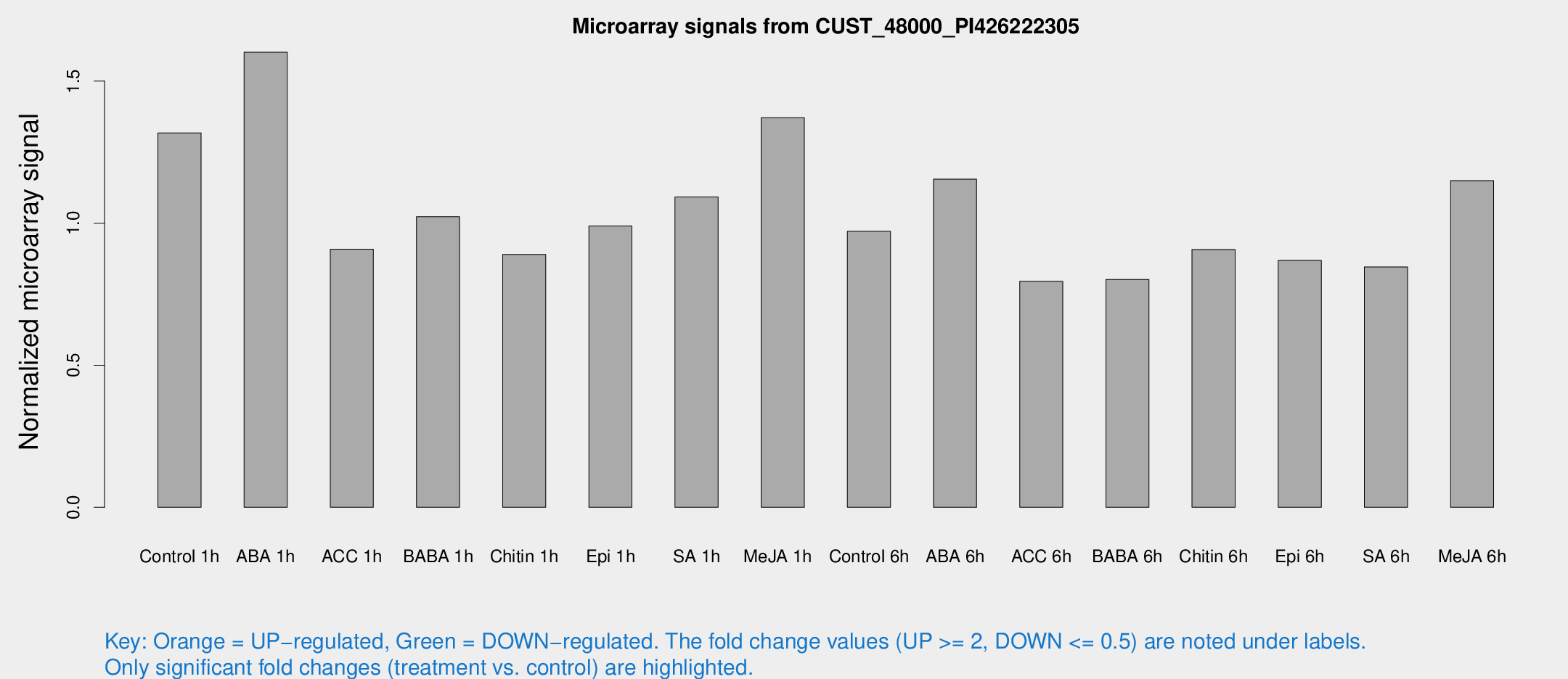
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 824.643 | 86.0286 | 1.31795 | 0.0762981 |
| ABA 1h | 888.979 | 100.528 | 1.60171 | 0.0926811 |
| ACC 1h | 652.158 | 192.109 | 0.908519 | 0.290337 |
| BABA 1h | 645.558 | 139.972 | 1.02291 | 0.148615 |
| Chitin 1h | 501.86 | 55.4029 | 0.889794 | 0.0517357 |
| Epi 1h | 534.27 | 44.9735 | 0.990172 | 0.102179 |
| SA 1h | 704.004 | 80.3097 | 1.09221 | 0.101783 |
| Me-JA 1h | 703.723 | 88.5562 | 1.37149 | 0.079491 |
| Control 6h | 649.119 | 164.884 | 0.971867 | 0.201215 |
| ABA 6h | 774.71 | 100.623 | 1.1552 | 0.120576 |
| ACC 6h | 575.16 | 80.3584 | 0.795243 | 0.0462554 |
| BABA 6h | 564.065 | 70.1796 | 0.802201 | 0.0971098 |
| Chitin 6h | 600.207 | 37.1041 | 0.907405 | 0.0527325 |
| Epi 6h | 614.932 | 73.394 | 0.868843 | 0.093899 |
| SA 6h | 566.09 | 158.448 | 0.845885 | 0.173712 |
| Me-JA 6h | 727.073 | 111.902 | 1.14948 | 0.0906264 |
Source Transcript PGSC0003DMT400074454 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G51050.1 | +3 | 5e-175 | 505 | 267/353 (76%) | Mitochondrial substrate carrier family protein | chr5:20753381-20755714 FORWARD LENGTH=487 |
