Probe CUST_48003_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48003_PI426222305 | JHI_St_60k_v1 | DMT400074453 | TTTCTTCCCCTTCTATTTGGATGGTGTAGCTTTCTGGTTTATCATGTGACATGCCGTTTT |
All Microarray Probes Designed to Gene DMG402028933
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48003_PI426222305 | JHI_St_60k_v1 | DMT400074453 | TTTCTTCCCCTTCTATTTGGATGGTGTAGCTTTCTGGTTTATCATGTGACATGCCGTTTT |
| CUST_48000_PI426222305 | JHI_St_60k_v1 | DMT400074454 | TTATCCTTTACAGGTTATTCGGACCAGAATGCAAGCTGATTCTGCATACAAAGGTATGGC |
Microarray Signals from CUST_48003_PI426222305
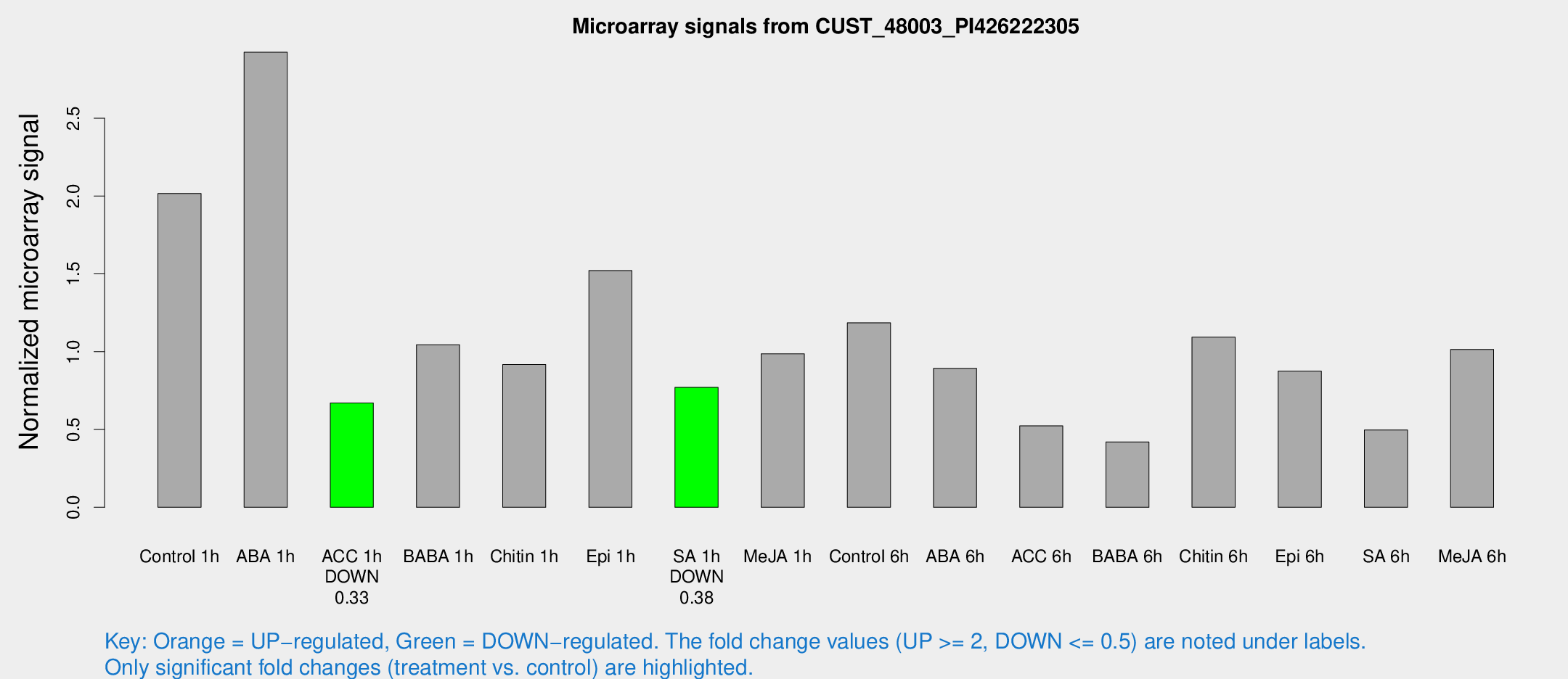
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 42.9791 | 9.19883 | 2.0171 | 0.285654 |
| ABA 1h | 53.172 | 4.47409 | 2.92483 | 0.246882 |
| ACC 1h | 14.2081 | 4.41772 | 0.670213 | 0.219505 |
| BABA 1h | 26.8101 | 10.2978 | 1.0452 | 0.603347 |
| Chitin 1h | 19.9938 | 7.07109 | 0.917369 | 0.599565 |
| Epi 1h | 27.4923 | 3.98313 | 1.52159 | 0.238417 |
| SA 1h | 16.1925 | 3.7741 | 0.771212 | 0.180849 |
| Me-JA 1h | 18.8348 | 7.2251 | 0.986108 | 0.315881 |
| Control 6h | 32.2465 | 12.6515 | 1.18585 | 0.756979 |
| ABA 6h | 24.4253 | 9.17845 | 0.892704 | 0.541881 |
| ACC 6h | 13.7516 | 4.57206 | 0.52339 | 0.21368 |
| BABA 6h | 9.94637 | 4.32752 | 0.420192 | 0.19526 |
| Chitin 6h | 24.0777 | 4.46979 | 1.09336 | 0.210047 |
| Epi 6h | 21.0701 | 4.98659 | 0.876182 | 0.232024 |
| SA 6h | 11.2625 | 4.22801 | 0.497209 | 0.260621 |
| Me-JA 6h | 22.809 | 7.02665 | 1.01475 | 0.250298 |
Source Transcript PGSC0003DMT400074453 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G51050.1 | +3 | 9e-146 | 432 | 231/300 (77%) | Mitochondrial substrate carrier family protein | chr5:20753381-20755714 FORWARD LENGTH=487 |
