Probe CUST_46837_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46837_PI426222305 | JHI_St_60k_v1 | DMT400001927 | CAGCTGTAATACTAGTGGCAGGTTCTTCCTTTATAATGTCCGTGCTTATATAAAAAACTT |
All Microarray Probes Designed to Gene DMG400000731
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46837_PI426222305 | JHI_St_60k_v1 | DMT400001927 | CAGCTGTAATACTAGTGGCAGGTTCTTCCTTTATAATGTCCGTGCTTATATAAAAAACTT |
| CUST_46811_PI426222305 | JHI_St_60k_v1 | DMT400001925 | CAATGTTCCTGTCCTCTTCTTCATTCAAGAGTTTATAAGTTGAGAGCTGGGGAAAGGGCA |
Microarray Signals from CUST_46837_PI426222305
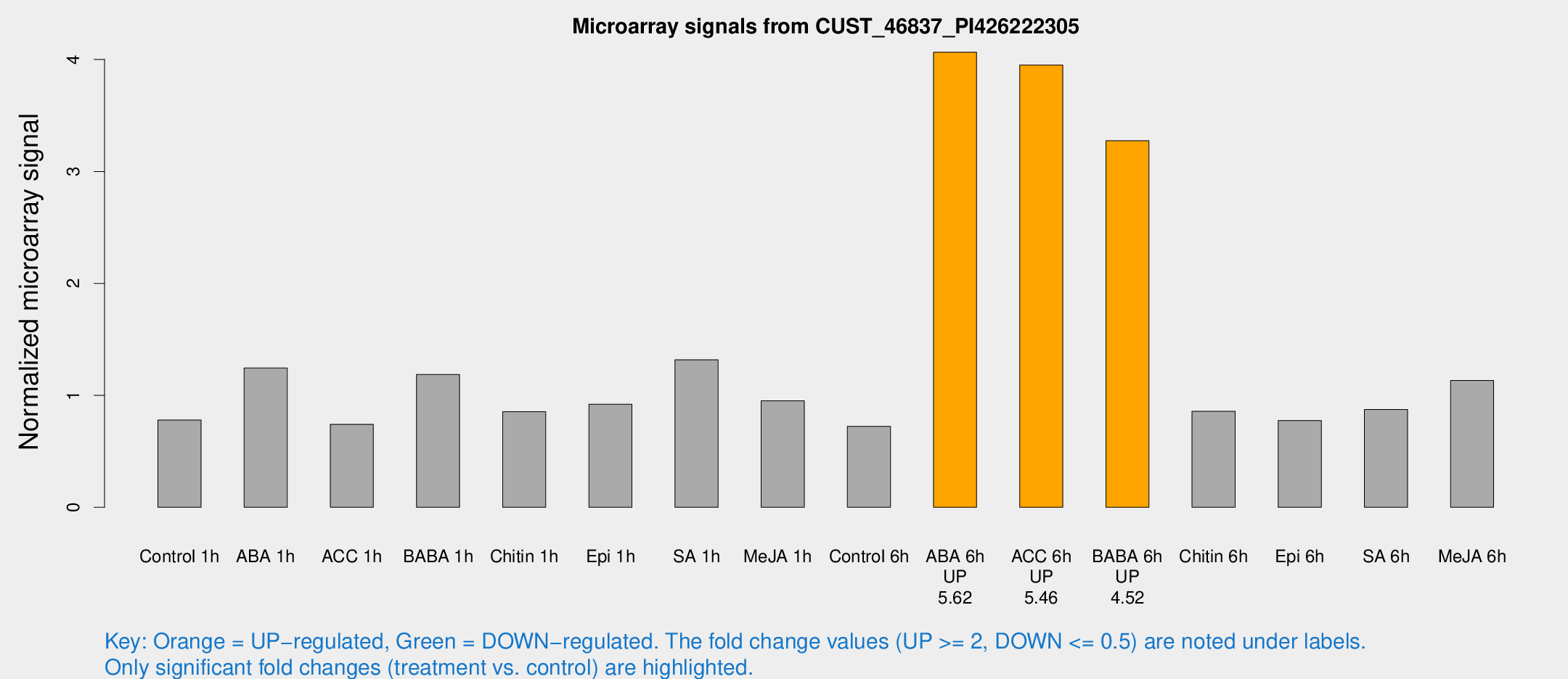
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 961.814 | 71.0809 | 0.779781 | 0.0451035 |
| ABA 1h | 1392.19 | 237.768 | 1.24391 | 0.191703 |
| ACC 1h | 1001.23 | 238.462 | 0.741643 | 0.153594 |
| BABA 1h | 1489.32 | 336.255 | 1.18687 | 0.18304 |
| Chitin 1h | 951.665 | 98.463 | 0.853823 | 0.0505289 |
| Epi 1h | 987.764 | 93.8134 | 0.921212 | 0.0835395 |
| SA 1h | 1669.19 | 123.923 | 1.31681 | 0.0760712 |
| Me-JA 1h | 971.14 | 136.57 | 0.951695 | 0.0605094 |
| Control 6h | 955.93 | 247.433 | 0.723773 | 0.149787 |
| ABA 6h | 5607.64 | 1205.41 | 4.06519 | 0.826607 |
| ACC 6h | 5652.74 | 721.614 | 3.95035 | 0.820731 |
| BABA 6h | 4762.31 | 1134.78 | 3.27386 | 0.780531 |
| Chitin 6h | 1124.17 | 69.2648 | 0.85752 | 0.0495952 |
| Epi 6h | 1121.84 | 235.517 | 0.775282 | 0.107562 |
| SA 6h | 1064.73 | 66.6945 | 0.873673 | 0.050901 |
| Me-JA 6h | 1385.06 | 80.0887 | 1.13282 | 0.100637 |
Source Transcript PGSC0003DMT400001927 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G21620.2 | +3 | 6e-86 | 259 | 134/186 (72%) | Adenine nucleotide alpha hydrolases-like superfamily protein | chr2:9248749-9249986 FORWARD LENGTH=193 |
