Probe CUST_46811_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46811_PI426222305 | JHI_St_60k_v1 | DMT400001925 | CAATGTTCCTGTCCTCTTCTTCATTCAAGAGTTTATAAGTTGAGAGCTGGGGAAAGGGCA |
All Microarray Probes Designed to Gene DMG400000731
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_46837_PI426222305 | JHI_St_60k_v1 | DMT400001927 | CAGCTGTAATACTAGTGGCAGGTTCTTCCTTTATAATGTCCGTGCTTATATAAAAAACTT |
| CUST_46811_PI426222305 | JHI_St_60k_v1 | DMT400001925 | CAATGTTCCTGTCCTCTTCTTCATTCAAGAGTTTATAAGTTGAGAGCTGGGGAAAGGGCA |
Microarray Signals from CUST_46811_PI426222305
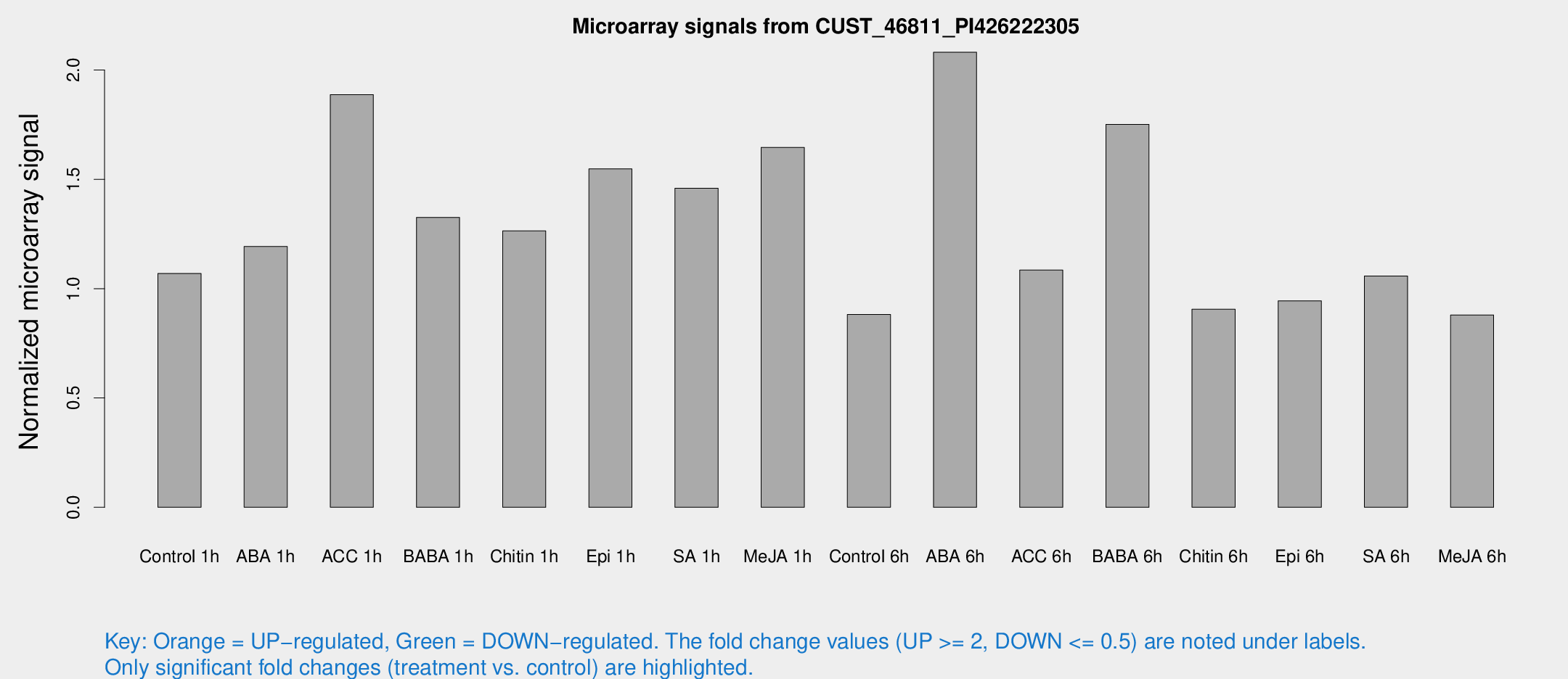
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8.29501 | 3.79848 | 1.06921 | 0.529444 |
| ABA 1h | 8.19757 | 3.65911 | 1.19284 | 0.582627 |
| ACC 1h | 28.1903 | 21.6436 | 1.88718 | 3.27215 |
| BABA 1h | 10.8788 | 4.08878 | 1.32598 | 0.702036 |
| Chitin 1h | 9.26941 | 3.81623 | 1.26398 | 0.609188 |
| Epi 1h | 11.0504 | 3.72975 | 1.54752 | 0.647145 |
| SA 1h | 13.1836 | 5.43172 | 1.45929 | 0.680684 |
| Me-JA 1h | 11.1618 | 3.88881 | 1.64615 | 0.691604 |
| Control 6h | 6.622 | 3.83737 | 0.881505 | 0.510659 |
| ABA 6h | 21.2354 | 10.184 | 2.08119 | 1.09721 |
| ACC 6h | 9.51443 | 4.83604 | 1.08492 | 0.551143 |
| BABA 6h | 14.9041 | 4.16788 | 1.75147 | 0.498275 |
| Chitin 6h | 7.23354 | 4.19265 | 0.90596 | 0.524573 |
| Epi 6h | 8.066 | 4.72749 | 0.94443 | 0.547368 |
| SA 6h | 7.95964 | 4.08628 | 1.05813 | 0.557544 |
| Me-JA 6h | 6.56953 | 3.81595 | 0.879646 | 0.510534 |
Source Transcript PGSC0003DMT400001925 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G21620.2 | +1 | 5e-65 | 201 | 109/157 (69%) | Adenine nucleotide alpha hydrolases-like superfamily protein | chr2:9248749-9249986 FORWARD LENGTH=193 |
