Probe CUST_45628_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45628_PI426222305 | JHI_St_60k_v1 | DMT400043551 | CTCTTGGTTTGACTTTTAACTATGGTTGACCTTGTGGTTTGTGTACAACTATACTTTGAT |
All Microarray Probes Designed to Gene DMG400016913
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45617_PI426222305 | JHI_St_60k_v1 | DMT400043552 | CTTGGTGAATTCTCAGTTATTTCAAAACCACTCCTCACAATAGAGAGGGGCTTGTCAAAT |
| CUST_45628_PI426222305 | JHI_St_60k_v1 | DMT400043551 | CTCTTGGTTTGACTTTTAACTATGGTTGACCTTGTGGTTTGTGTACAACTATACTTTGAT |
Microarray Signals from CUST_45628_PI426222305
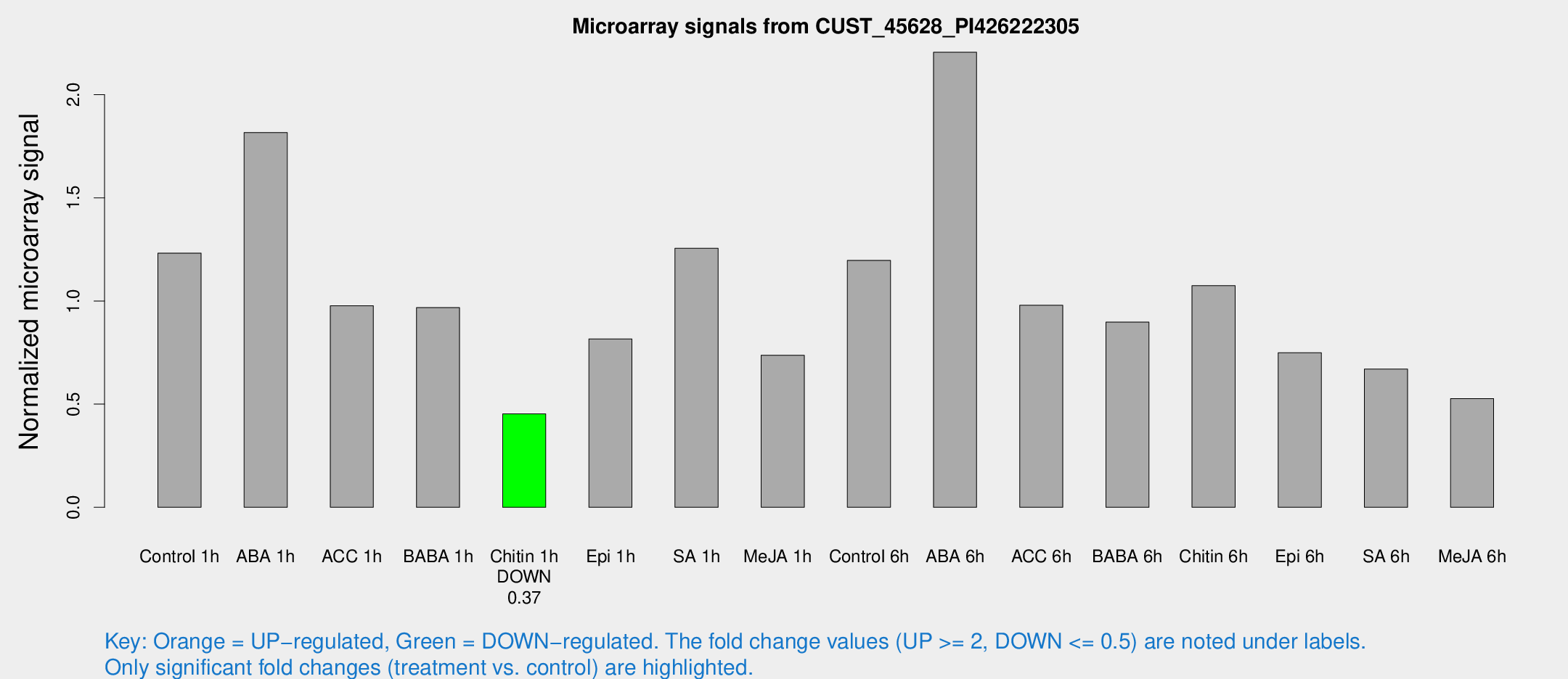
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18.4246 | 3.70772 | 1.23213 | 0.250741 |
| ABA 1h | 24.5159 | 3.86546 | 1.81655 | 0.296843 |
| ACC 1h | 15.5781 | 4.54474 | 0.976816 | 0.343995 |
| BABA 1h | 16.107 | 5.52729 | 0.967786 | 0.338432 |
| Chitin 1h | 6.06054 | 3.52652 | 0.452845 | 0.262475 |
| Epi 1h | 14.275 | 8.08954 | 0.81611 | 0.518151 |
| SA 1h | 20.7833 | 6.29685 | 1.25573 | 0.387488 |
| Me-JA 1h | 9.94413 | 3.73814 | 0.73726 | 0.347119 |
| Control 6h | 23.7486 | 11.7642 | 1.19679 | 0.704318 |
| ABA 6h | 34.9953 | 4.30418 | 2.20565 | 0.275115 |
| ACC 6h | 17.2832 | 4.58266 | 0.979434 | 0.268119 |
| BABA 6h | 18.1801 | 8.23346 | 0.897483 | 0.433703 |
| Chitin 6h | 19.8868 | 6.42329 | 1.07419 | 0.506387 |
| Epi 6h | 16.0615 | 8.16756 | 0.749475 | 0.414327 |
| SA 6h | 11.124 | 4.1256 | 0.670009 | 0.312682 |
| Me-JA 6h | 8.16736 | 3.71939 | 0.526784 | 0.262474 |
Source Transcript PGSC0003DMT400043551 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G10540.1 | +3 | 0.0 | 611 | 309/380 (81%) | 3-phosphoinositide-dependent protein kinase | chr3:3289916-3292429 FORWARD LENGTH=486 |
