Probe CUST_45617_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45617_PI426222305 | JHI_St_60k_v1 | DMT400043552 | CTTGGTGAATTCTCAGTTATTTCAAAACCACTCCTCACAATAGAGAGGGGCTTGTCAAAT |
All Microarray Probes Designed to Gene DMG400016913
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45617_PI426222305 | JHI_St_60k_v1 | DMT400043552 | CTTGGTGAATTCTCAGTTATTTCAAAACCACTCCTCACAATAGAGAGGGGCTTGTCAAAT |
| CUST_45628_PI426222305 | JHI_St_60k_v1 | DMT400043551 | CTCTTGGTTTGACTTTTAACTATGGTTGACCTTGTGGTTTGTGTACAACTATACTTTGAT |
Microarray Signals from CUST_45617_PI426222305
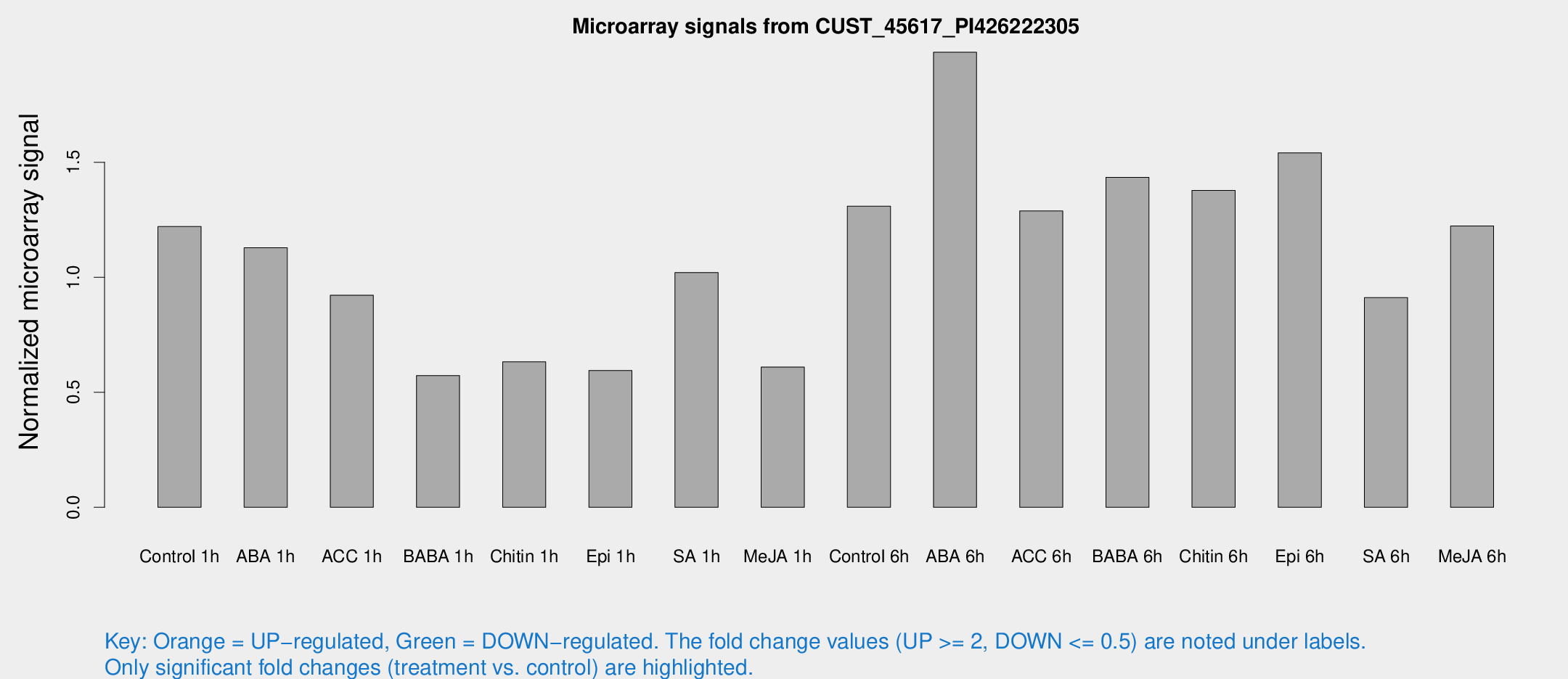
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 24.8077 | 7.83301 | 1.22039 | 0.294504 |
| ABA 1h | 20.0569 | 5.75877 | 1.12866 | 0.396181 |
| ACC 1h | 21.0848 | 7.33753 | 0.921995 | 0.401101 |
| BABA 1h | 10.5007 | 3.27034 | 0.573041 | 0.187755 |
| Chitin 1h | 11.3465 | 3.1407 | 0.632599 | 0.210064 |
| Epi 1h | 10.2583 | 3.08289 | 0.594794 | 0.20596 |
| SA 1h | 21.0806 | 6.20132 | 1.02067 | 0.290036 |
| Me-JA 1h | 9.63474 | 3.19296 | 0.609503 | 0.231551 |
| Control 6h | 24.8578 | 3.48031 | 1.30885 | 0.193248 |
| ABA 6h | 39.5553 | 4.47449 | 1.97848 | 0.32053 |
| ACC 6h | 27.5482 | 4.2045 | 1.28846 | 0.206213 |
| BABA 6h | 30.2581 | 3.93717 | 1.43465 | 0.190401 |
| Chitin 6h | 28.764 | 6.78179 | 1.37734 | 0.318095 |
| Epi 6h | 33.2064 | 5.40042 | 1.54128 | 0.350748 |
| SA 6h | 17.2074 | 3.46719 | 0.91166 | 0.196968 |
| Me-JA 6h | 24.8641 | 7.48801 | 1.22316 | 0.347283 |
Source Transcript PGSC0003DMT400043552 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G10540.1 | +3 | 0.0 | 781 | 397/481 (83%) | 3-phosphoinositide-dependent protein kinase | chr3:3289916-3292429 FORWARD LENGTH=486 |
