Probe CUST_44977_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44977_PI426222305 | JHI_St_60k_v1 | DMT400011224 | GGTCTTTTATTTCCCTGCATTCAATCTTTCAATTGCAGTGTCCAACTTTTGGTCTATGTA |
All Microarray Probes Designed to Gene DMG400004396
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44964_PI426222305 | JHI_St_60k_v1 | DMT400011223 | TGGTTCTTCAAATAACACAGCAGCTTCTGCAGCATCTTCGCCTTCCGCATCACTTTTGTA |
| CUST_44977_PI426222305 | JHI_St_60k_v1 | DMT400011224 | GGTCTTTTATTTCCCTGCATTCAATCTTTCAATTGCAGTGTCCAACTTTTGGTCTATGTA |
Microarray Signals from CUST_44977_PI426222305
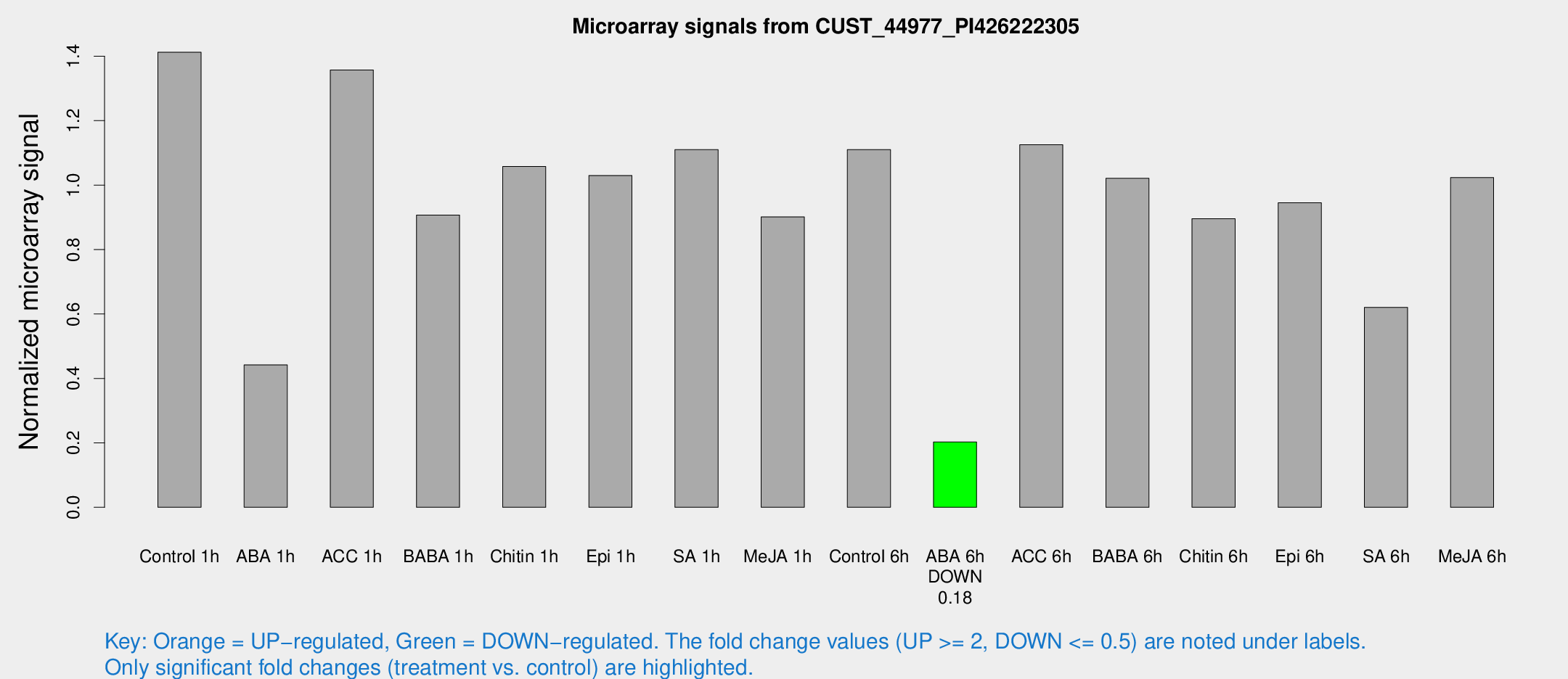
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 341.244 | 44.1621 | 1.41246 | 0.128261 |
| ABA 1h | 94.2834 | 11.4826 | 0.441902 | 0.0772769 |
| ACC 1h | 338.522 | 46.648 | 1.35707 | 0.0986041 |
| BABA 1h | 219.018 | 47.4482 | 0.907041 | 0.12707 |
| Chitin 1h | 236.389 | 49.8256 | 1.05776 | 0.284579 |
| Epi 1h | 211.982 | 12.6927 | 1.02991 | 0.061668 |
| SA 1h | 273.677 | 25.7048 | 1.10996 | 0.0655817 |
| Me-JA 1h | 176.564 | 16.5246 | 0.901097 | 0.0551745 |
| Control 6h | 283.971 | 66.6939 | 1.11032 | 0.211545 |
| ABA 6h | 53.2276 | 10.6706 | 0.202674 | 0.0326107 |
| ACC 6h | 324.798 | 80.6883 | 1.12542 | 0.121932 |
| BABA 6h | 273.443 | 21.7833 | 1.02113 | 0.0608999 |
| Chitin 6h | 227.522 | 13.8179 | 0.895704 | 0.0542387 |
| Epi 6h | 268.672 | 65.2558 | 0.945035 | 0.277083 |
| SA 6h | 151.185 | 26.2124 | 0.620542 | 0.0464634 |
| Me-JA 6h | 246.936 | 33.8428 | 1.02326 | 0.113149 |
Source Transcript PGSC0003DMT400011224 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G10530.1 | +1 | 4e-149 | 455 | 290/646 (45%) | Concanavalin A-like lectin protein kinase family protein | chr5:3324978-3326933 REVERSE LENGTH=651 |
