Probe CUST_44964_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44964_PI426222305 | JHI_St_60k_v1 | DMT400011223 | TGGTTCTTCAAATAACACAGCAGCTTCTGCAGCATCTTCGCCTTCCGCATCACTTTTGTA |
All Microarray Probes Designed to Gene DMG400004396
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44964_PI426222305 | JHI_St_60k_v1 | DMT400011223 | TGGTTCTTCAAATAACACAGCAGCTTCTGCAGCATCTTCGCCTTCCGCATCACTTTTGTA |
| CUST_44977_PI426222305 | JHI_St_60k_v1 | DMT400011224 | GGTCTTTTATTTCCCTGCATTCAATCTTTCAATTGCAGTGTCCAACTTTTGGTCTATGTA |
Microarray Signals from CUST_44964_PI426222305
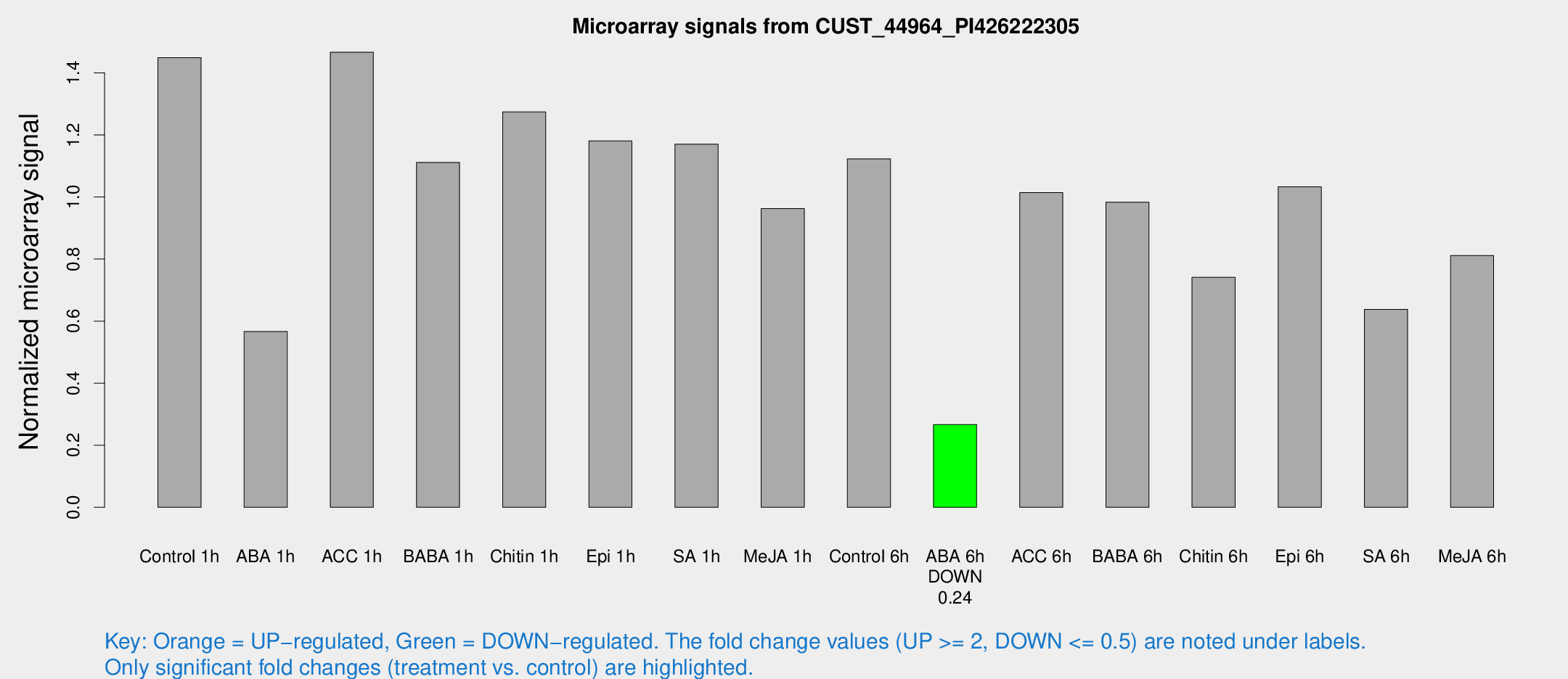
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 389.785 | 45.7564 | 1.44858 | 0.128147 |
| ABA 1h | 138.632 | 28.8792 | 0.566293 | 0.161352 |
| ACC 1h | 402.007 | 29.6396 | 1.46638 | 0.0861324 |
| BABA 1h | 294.176 | 53.7379 | 1.11118 | 0.118127 |
| Chitin 1h | 320.134 | 74.6108 | 1.2739 | 0.344978 |
| Epi 1h | 271.689 | 16.1277 | 1.18085 | 0.070013 |
| SA 1h | 328.461 | 52.7706 | 1.17022 | 0.15911 |
| Me-JA 1h | 211.886 | 25.5694 | 0.962506 | 0.0583087 |
| Control 6h | 314.486 | 64.434 | 1.12257 | 0.162107 |
| ABA 6h | 77.0997 | 11.5407 | 0.26673 | 0.0275604 |
| ACC 6h | 318.256 | 57.4126 | 1.01391 | 0.0603773 |
| BABA 6h | 294.644 | 26.2321 | 0.983257 | 0.116842 |
| Chitin 6h | 212.922 | 26.1734 | 0.740907 | 0.0783839 |
| Epi 6h | 324.29 | 72.5779 | 1.0328 | 0.262676 |
| SA 6h | 168.928 | 15.0927 | 0.638012 | 0.0400874 |
| Me-JA 6h | 220.777 | 37.0462 | 0.811494 | 0.184034 |
Source Transcript PGSC0003DMT400011223 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G10530.1 | +1 | 3e-122 | 377 | 226/484 (47%) | Concanavalin A-like lectin protein kinase family protein | chr5:3324978-3326933 REVERSE LENGTH=651 |
