Probe CUST_44691_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44691_PI426222305 | JHI_St_60k_v1 | DMT400045281 | ACTAAAGTCATGATGAGTTGGACAGTTGGTCAGCAACAAGCCAACAACAACTTTTTTCAG |
All Microarray Probes Designed to Gene DMG400017564
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44727_PI426222305 | JHI_St_60k_v1 | DMT400045282 | CAGGATTTGTACTTTACACGGTTCGAGTTCGGAATTCTATCGTGATTCATTGAGTTCAAA |
| CUST_44691_PI426222305 | JHI_St_60k_v1 | DMT400045281 | ACTAAAGTCATGATGAGTTGGACAGTTGGTCAGCAACAAGCCAACAACAACTTTTTTCAG |
Microarray Signals from CUST_44691_PI426222305
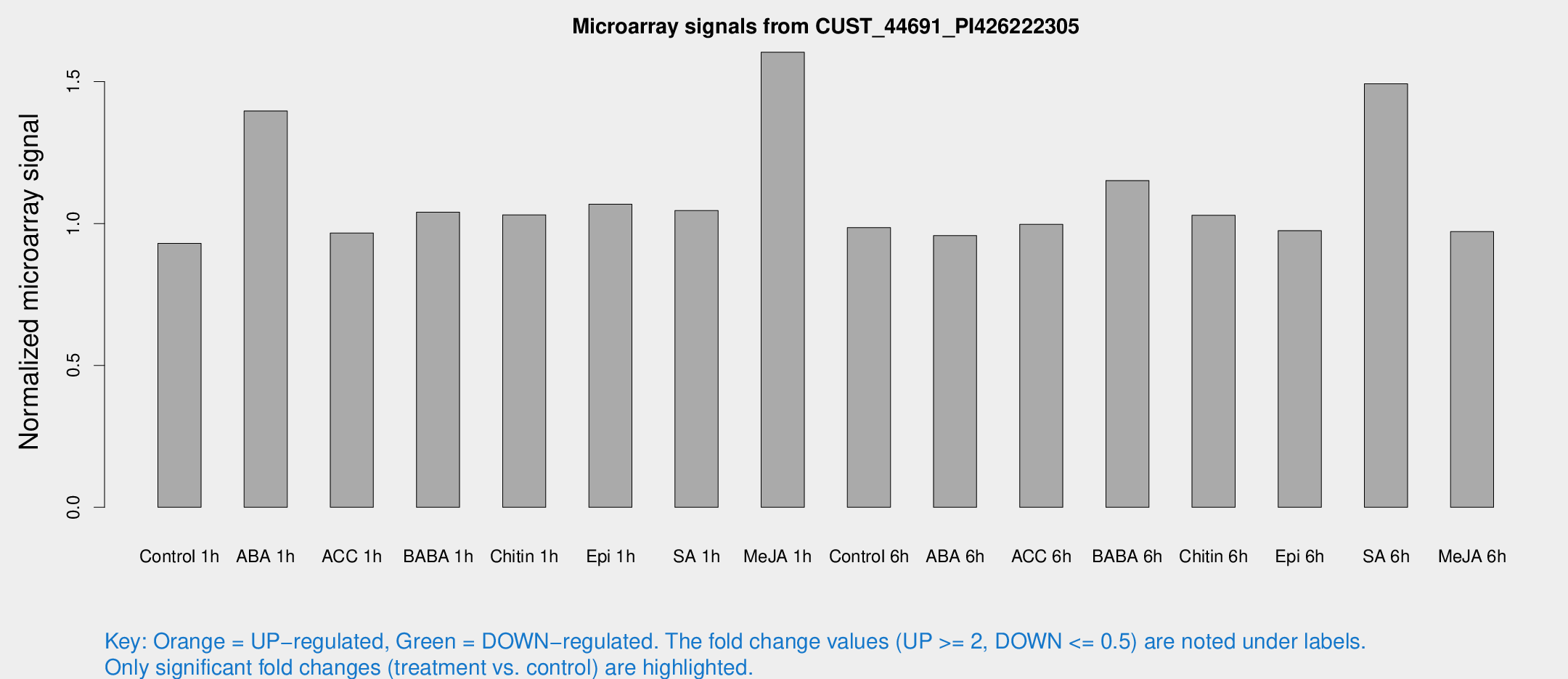
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.59488 | 3.24067 | 0.93007 | 0.538559 |
| ABA 1h | 8.41676 | 3.12615 | 1.39685 | 0.657432 |
| ACC 1h | 5.9712 | 3.46025 | 0.966407 | 0.559785 |
| BABA 1h | 6.03241 | 3.49551 | 1.04005 | 0.60231 |
| Chitin 1h | 5.588 | 3.2454 | 1.03062 | 0.597069 |
| Epi 1h | 5.57412 | 3.23825 | 1.06798 | 0.618699 |
| SA 1h | 6.52095 | 3.2484 | 1.04561 | 0.529423 |
| Me-JA 1h | 8.57219 | 3.3993 | 1.60376 | 0.760433 |
| Control 6h | 5.94092 | 3.442 | 0.985517 | 0.570919 |
| ABA 6h | 6.12819 | 3.55391 | 0.957319 | 0.554549 |
| ACC 6h | 7.03227 | 4.16869 | 0.997075 | 0.577532 |
| BABA 6h | 7.99142 | 3.80379 | 1.15152 | 0.581115 |
| Chitin 6h | 6.58834 | 3.81646 | 1.02889 | 0.595871 |
| Epi 6h | 6.66087 | 3.8888 | 0.975117 | 0.565198 |
| SA 6h | 9.15046 | 3.59222 | 1.49274 | 0.630596 |
| Me-JA 6h | 5.82145 | 3.37332 | 0.97152 | 0.562649 |
Source Transcript PGSC0003DMT400045281 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G18690.1 | +1 | 3e-17 | 77 | 62/138 (45%) | MAP kinase substrate 1 | chr3:6429755-6430423 REVERSE LENGTH=222 |
