Probe CUST_44727_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44727_PI426222305 | JHI_St_60k_v1 | DMT400045282 | CAGGATTTGTACTTTACACGGTTCGAGTTCGGAATTCTATCGTGATTCATTGAGTTCAAA |
All Microarray Probes Designed to Gene DMG400017564
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44727_PI426222305 | JHI_St_60k_v1 | DMT400045282 | CAGGATTTGTACTTTACACGGTTCGAGTTCGGAATTCTATCGTGATTCATTGAGTTCAAA |
| CUST_44691_PI426222305 | JHI_St_60k_v1 | DMT400045281 | ACTAAAGTCATGATGAGTTGGACAGTTGGTCAGCAACAAGCCAACAACAACTTTTTTCAG |
Microarray Signals from CUST_44727_PI426222305
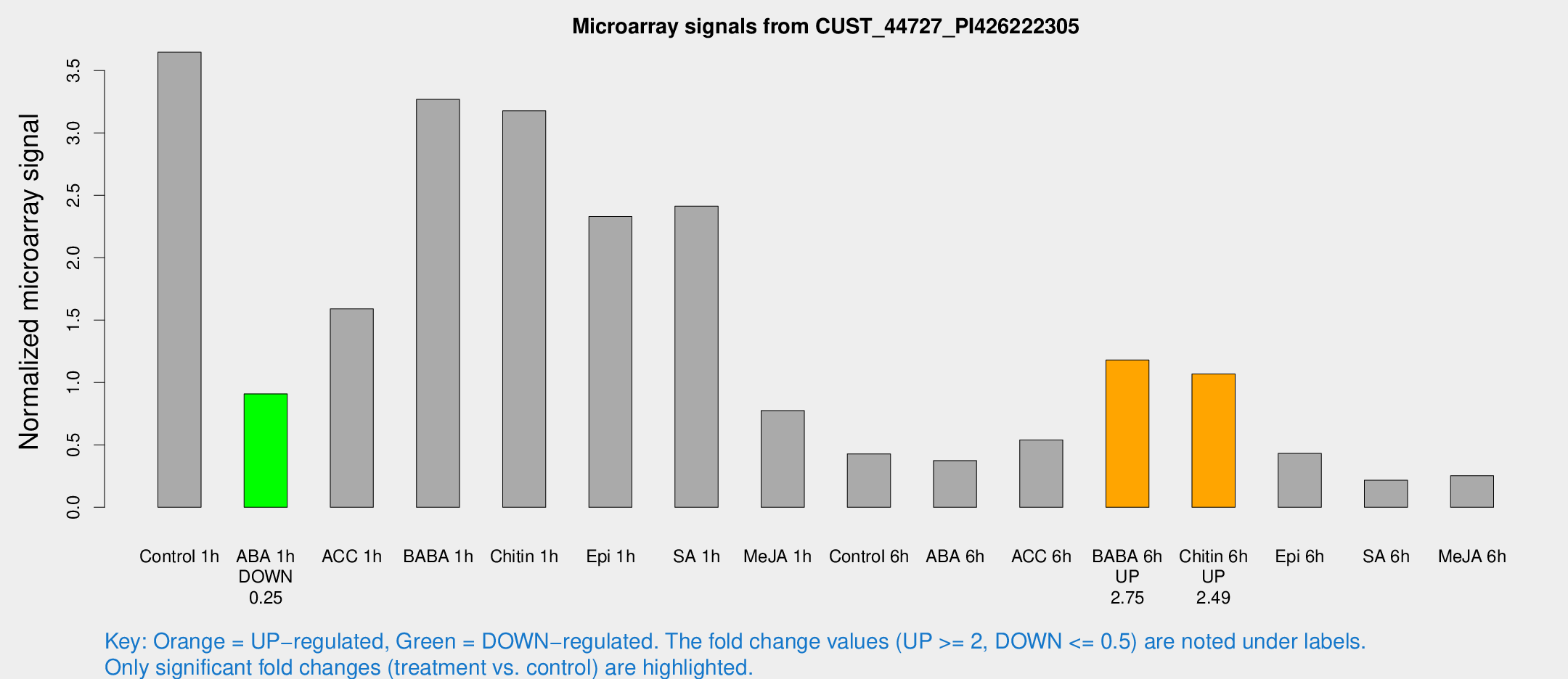
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 193.044 | 35.2784 | 3.6472 | 0.486726 |
| ABA 1h | 41.3244 | 4.04373 | 0.909398 | 0.0893507 |
| ACC 1h | 122.965 | 54.5932 | 1.59005 | 1.44609 |
| BABA 1h | 162.784 | 13.7816 | 3.26939 | 0.245907 |
| Chitin 1h | 146.989 | 9.16121 | 3.17716 | 0.399266 |
| Epi 1h | 104.363 | 9.9269 | 2.33006 | 0.164269 |
| SA 1h | 133.48 | 27.273 | 2.41258 | 0.565544 |
| Me-JA 1h | 41.0278 | 16.2185 | 0.774675 | 0.501989 |
| Control 6h | 22.9151 | 4.28179 | 0.428484 | 0.0799499 |
| ABA 6h | 20.5965 | 3.81161 | 0.374476 | 0.0704626 |
| ACC 6h | 31.9783 | 4.51265 | 0.540327 | 0.11574 |
| BABA 6h | 71.6252 | 15.9844 | 1.17999 | 0.244952 |
| Chitin 6h | 58.435 | 5.14511 | 1.06888 | 0.0942939 |
| Epi 6h | 26.6117 | 5.94108 | 0.432071 | 0.0869763 |
| SA 6h | 11.6365 | 3.81222 | 0.216523 | 0.0811834 |
| Me-JA 6h | 15.5011 | 6.2721 | 0.253053 | 0.0981604 |
Source Transcript PGSC0003DMT400045282 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G18690.1 | +3 | 3e-22 | 93 | 91/251 (36%) | MAP kinase substrate 1 | chr3:6429755-6430423 REVERSE LENGTH=222 |
