Probe CUST_44355_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44355_PI426222305 | JHI_St_60k_v1 | DMT400010084 | GATTTAGGTCTCGTATTTCAGCTACAAGTCAATCGTGTGTCATTAATTCATATTTCTTCC |
All Microarray Probes Designed to Gene DMG400003948
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44349_PI426222305 | JHI_St_60k_v1 | DMT400010083 | AAAGAAGAGATTTGTAAGGCTAATCCTGTAGTTAAGGCTGATAAAATCCAGCGGTTTTGA |
| CUST_44355_PI426222305 | JHI_St_60k_v1 | DMT400010084 | GATTTAGGTCTCGTATTTCAGCTACAAGTCAATCGTGTGTCATTAATTCATATTTCTTCC |
Microarray Signals from CUST_44355_PI426222305
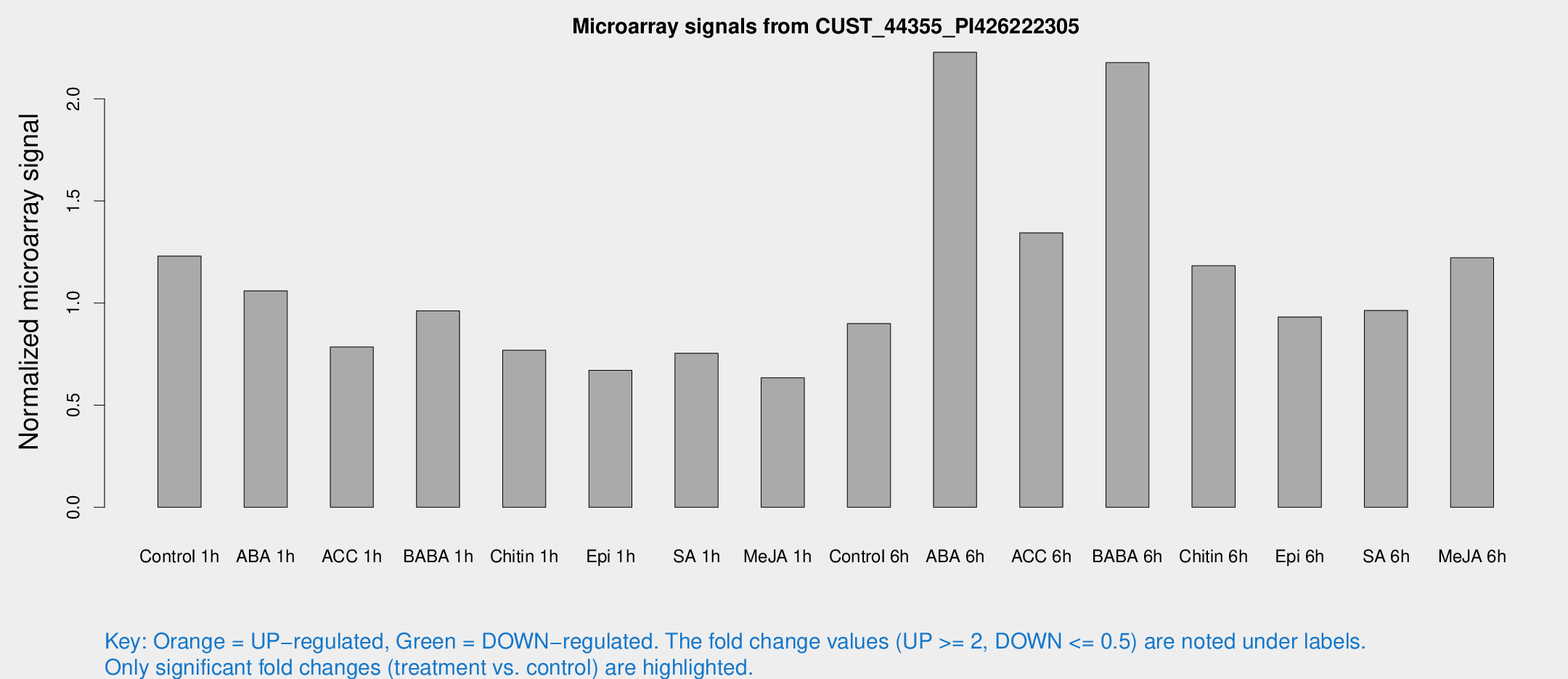
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 90.0083 | 6.93148 | 1.22998 | 0.0848115 |
| ABA 1h | 77.3099 | 28.7885 | 1.0599 | 0.31406 |
| ACC 1h | 63.0035 | 15.8768 | 0.785215 | 0.168373 |
| BABA 1h | 70.0125 | 12.4988 | 0.962416 | 0.0913942 |
| Chitin 1h | 51.7601 | 9.04683 | 0.769127 | 0.082697 |
| Epi 1h | 44.8129 | 11.3552 | 0.670322 | 0.149681 |
| SA 1h | 59.9797 | 13.5582 | 0.753838 | 0.158612 |
| Me-JA 1h | 49.6426 | 24.4934 | 0.63424 | 0.339287 |
| Control 6h | 74.4148 | 26.2949 | 0.899538 | 0.281603 |
| ABA 6h | 194.181 | 56.5778 | 2.22829 | 0.800887 |
| ACC 6h | 114.04 | 15.2607 | 1.34353 | 0.0905177 |
| BABA 6h | 198.934 | 63.2098 | 2.17757 | 0.725816 |
| Chitin 6h | 92.0844 | 6.58667 | 1.18276 | 0.102749 |
| Epi 6h | 78.4861 | 13.1 | 0.932182 | 0.148732 |
| SA 6h | 72.6257 | 14.7853 | 0.964038 | 0.336394 |
| Me-JA 6h | 106.667 | 47.1075 | 1.22209 | 0.460384 |
Source Transcript PGSC0003DMT400010084 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G12040.1 | +3 | 4e-49 | 167 | 85/174 (49%) | A20/AN1-like zinc finger family protein | chr4:7215341-7215868 FORWARD LENGTH=175 |
