Probe CUST_44349_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44349_PI426222305 | JHI_St_60k_v1 | DMT400010083 | AAAGAAGAGATTTGTAAGGCTAATCCTGTAGTTAAGGCTGATAAAATCCAGCGGTTTTGA |
All Microarray Probes Designed to Gene DMG400003948
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44349_PI426222305 | JHI_St_60k_v1 | DMT400010083 | AAAGAAGAGATTTGTAAGGCTAATCCTGTAGTTAAGGCTGATAAAATCCAGCGGTTTTGA |
| CUST_44355_PI426222305 | JHI_St_60k_v1 | DMT400010084 | GATTTAGGTCTCGTATTTCAGCTACAAGTCAATCGTGTGTCATTAATTCATATTTCTTCC |
Microarray Signals from CUST_44349_PI426222305
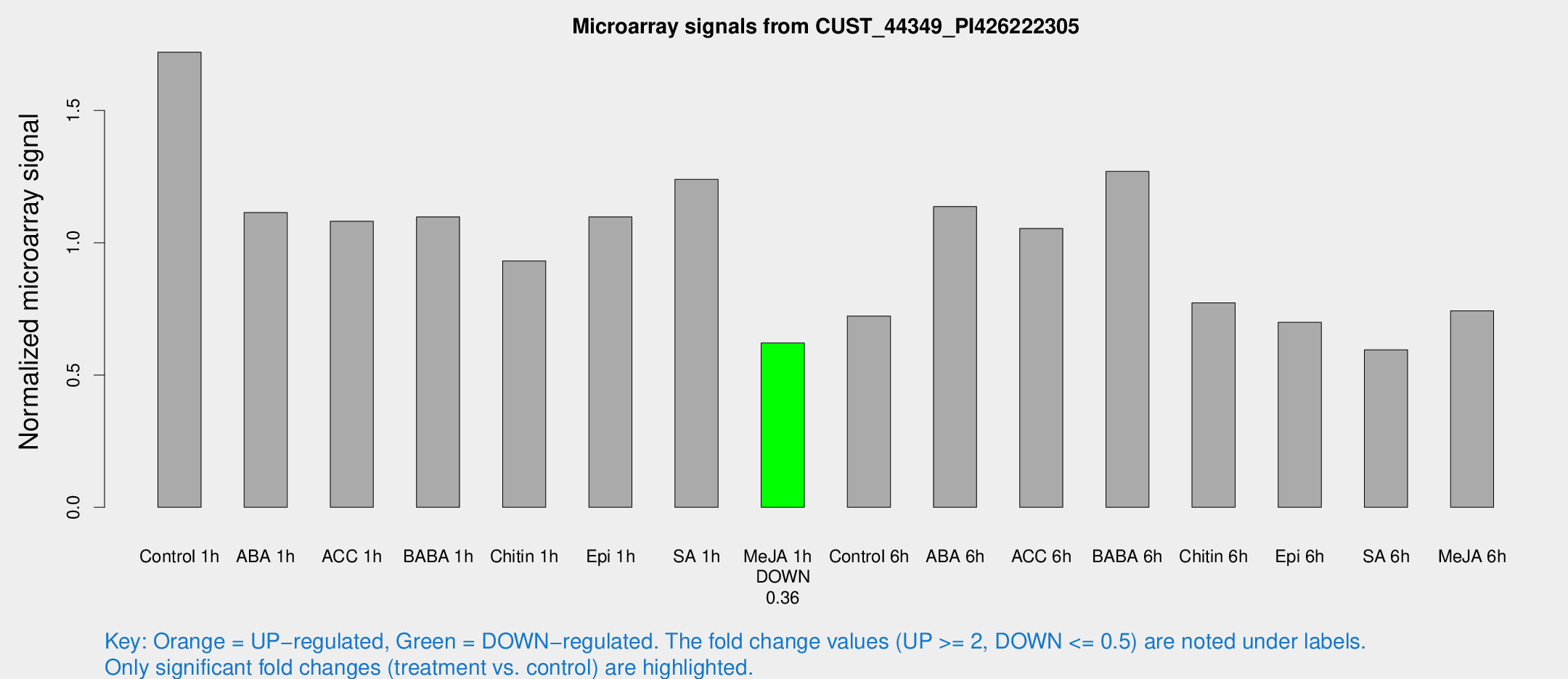
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 13449.7 | 778.615 | 1.7202 | 0.0993166 |
| ABA 1h | 7765.77 | 797.876 | 1.11421 | 0.0643309 |
| ACC 1h | 9782.78 | 2906.27 | 1.08128 | 0.346917 |
| BABA 1h | 8902.81 | 2266.2 | 1.09792 | 0.210351 |
| Chitin 1h | 6684.71 | 1040.79 | 0.931194 | 0.11307 |
| Epi 1h | 7413.09 | 428.246 | 1.0978 | 0.0633837 |
| SA 1h | 10413.7 | 2247.24 | 1.23947 | 0.198792 |
| Me-JA 1h | 4054.46 | 664.347 | 0.621298 | 0.0571763 |
| Control 6h | 5683.05 | 427.209 | 0.723093 | 0.0446647 |
| ABA 6h | 9610.92 | 1375.88 | 1.137 | 0.0960205 |
| ACC 6h | 9486.35 | 734.822 | 1.05399 | 0.169534 |
| BABA 6h | 11204.4 | 1198.12 | 1.26962 | 0.113692 |
| Chitin 6h | 6428.28 | 371.801 | 0.773151 | 0.0446405 |
| Epi 6h | 6277.66 | 879.065 | 0.699352 | 0.0486874 |
| SA 6h | 5283.91 | 1632.02 | 0.595454 | 0.182739 |
| Me-JA 6h | 5972.06 | 1147.76 | 0.742684 | 0.112777 |
Source Transcript PGSC0003DMT400010083 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G22820.1 | +1 | 5e-34 | 119 | 65/131 (50%) | A20/AN1-like zinc finger family protein | chr4:11987871-11988401 REVERSE LENGTH=176 |
