Probe CUST_44305_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44305_PI426222305 | JHI_St_60k_v1 | DMT400010101 | GCTTATCTTTCGAGGTTCTGTCAAAAGCATTCACTCTACTTGTGAAATTAGACTTTTTGA |
All Microarray Probes Designed to Gene DMG400003957
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44287_PI426222305 | JHI_St_60k_v1 | DMT400010100 | GCTTATCTTTCGAGGTTCTGTCAAAAGCATTCACTCTACTTGTGAAATTAGACTTTTTGA |
| CUST_44305_PI426222305 | JHI_St_60k_v1 | DMT400010101 | GCTTATCTTTCGAGGTTCTGTCAAAAGCATTCACTCTACTTGTGAAATTAGACTTTTTGA |
Microarray Signals from CUST_44305_PI426222305
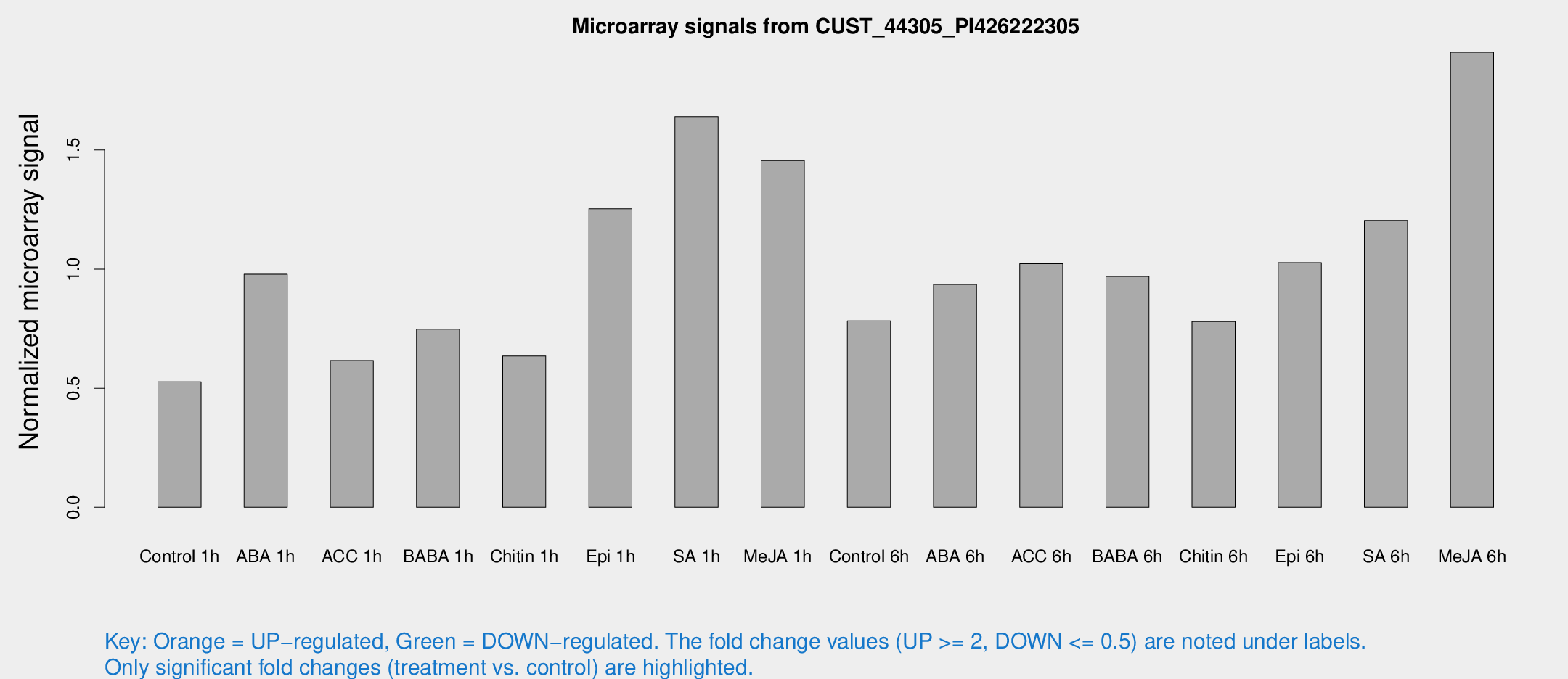
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.3474 | 2.95893 | 0.52695 | 0.251053 |
| ABA 1h | 12.778 | 3.90942 | 0.978924 | 0.533644 |
| ACC 1h | 9.08833 | 3.41287 | 0.616433 | 0.288757 |
| BABA 1h | 11.1696 | 4.86863 | 0.748258 | 0.297707 |
| Chitin 1h | 7.67169 | 2.97083 | 0.635625 | 0.268706 |
| Epi 1h | 14.4175 | 3.02643 | 1.25354 | 0.283353 |
| SA 1h | 22.163 | 3.23507 | 1.64022 | 0.250004 |
| Me-JA 1h | 17.391 | 5.14383 | 1.45644 | 0.476979 |
| Control 6h | 10.318 | 3.07406 | 0.782737 | 0.244943 |
| ABA 6h | 15.5003 | 5.51658 | 0.93642 | 0.471672 |
| ACC 6h | 16.2674 | 3.89707 | 1.0229 | 0.47781 |
| BABA 6h | 16.7762 | 5.69545 | 0.969748 | 0.479085 |
| Chitin 6h | 10.8248 | 3.41354 | 0.77971 | 0.251987 |
| Epi 6h | 16.1783 | 4.73391 | 1.02726 | 0.285354 |
| SA 6h | 16.75 | 4.23445 | 1.20455 | 0.312082 |
| Me-JA 6h | 24.7821 | 3.35522 | 1.91067 | 0.282182 |
Source Transcript PGSC0003DMT400010101 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G45360.1 | +3 | 2e-79 | 244 | 130/220 (59%) | Protein of unknown function (DUF1442) | chr2:18698619-18699360 FORWARD LENGTH=215 |
