Probe CUST_44287_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44287_PI426222305 | JHI_St_60k_v1 | DMT400010100 | GCTTATCTTTCGAGGTTCTGTCAAAAGCATTCACTCTACTTGTGAAATTAGACTTTTTGA |
All Microarray Probes Designed to Gene DMG400003957
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44287_PI426222305 | JHI_St_60k_v1 | DMT400010100 | GCTTATCTTTCGAGGTTCTGTCAAAAGCATTCACTCTACTTGTGAAATTAGACTTTTTGA |
| CUST_44305_PI426222305 | JHI_St_60k_v1 | DMT400010101 | GCTTATCTTTCGAGGTTCTGTCAAAAGCATTCACTCTACTTGTGAAATTAGACTTTTTGA |
Microarray Signals from CUST_44287_PI426222305
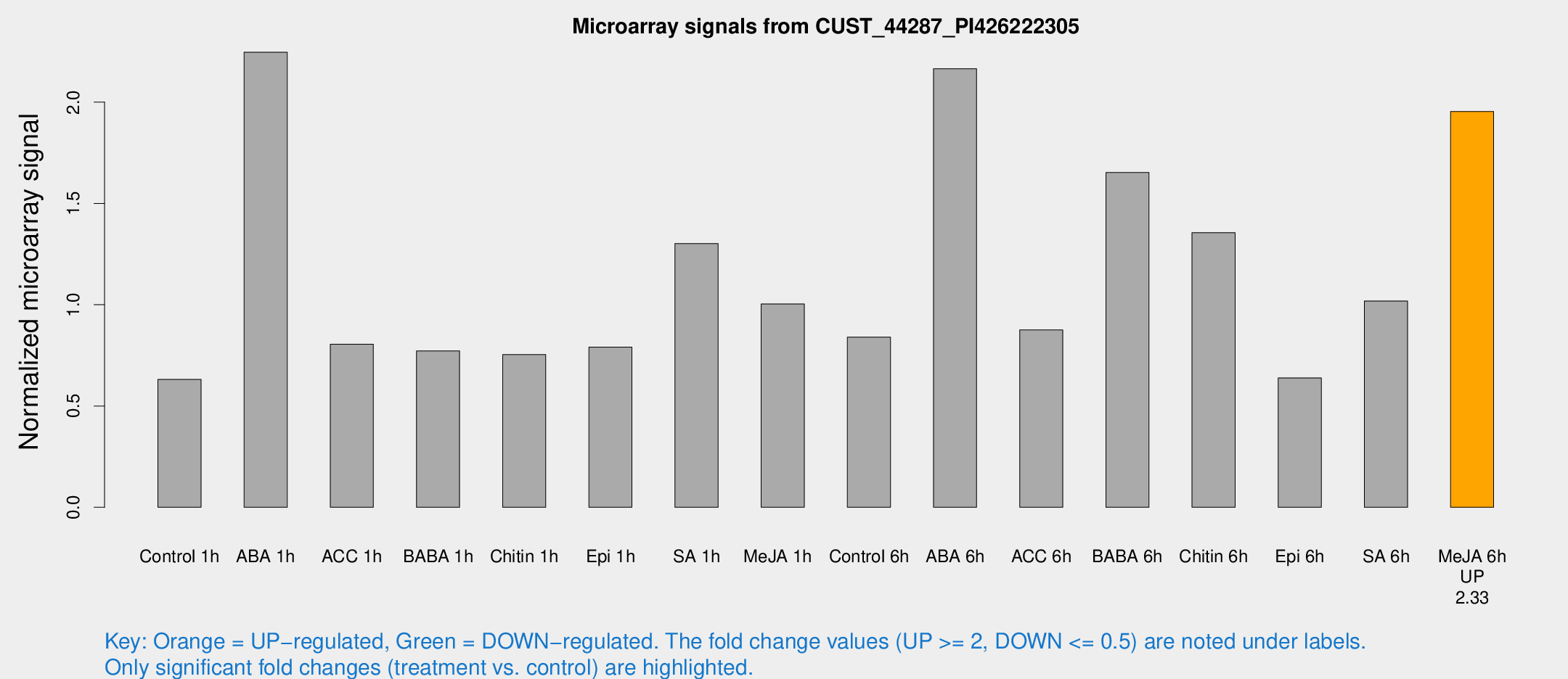
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.63399 | 3.72354 | 0.631082 | 0.321644 |
| ABA 1h | 26.5831 | 8.55713 | 2.24657 | 0.690347 |
| ACC 1h | 10.5808 | 4.33521 | 0.804866 | 0.382025 |
| BABA 1h | 9.14325 | 3.88158 | 0.772137 | 0.351221 |
| Chitin 1h | 8.6404 | 3.67102 | 0.753736 | 0.365406 |
| Epi 1h | 8.31565 | 3.62489 | 0.790572 | 0.364551 |
| SA 1h | 19.0797 | 6.47548 | 1.30163 | 0.631713 |
| Me-JA 1h | 10.5395 | 3.81648 | 1.00348 | 0.417886 |
| Control 6h | 10.3925 | 3.92204 | 0.840304 | 0.349361 |
| ABA 6h | 30.0439 | 7.94382 | 2.16545 | 0.656088 |
| ACC 6h | 12.0702 | 4.6698 | 0.875851 | 0.354213 |
| BABA 6h | 22.0058 | 4.39626 | 1.65273 | 0.333106 |
| Chitin 6h | 20.7718 | 7.95223 | 1.35607 | 0.733804 |
| Epi 6h | 8.69799 | 4.81028 | 0.638698 | 0.34779 |
| SA 6h | 12.9531 | 4.20308 | 1.01867 | 0.384678 |
| Me-JA 6h | 23.6269 | 3.93931 | 1.95423 | 0.451375 |
Source Transcript PGSC0003DMT400010100 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G45360.1 | +1 | 5e-66 | 211 | 113/207 (55%) | Protein of unknown function (DUF1442) | chr2:18698619-18699360 FORWARD LENGTH=215 |
