Probe CUST_44204_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44204_PI426222305 | JHI_St_60k_v1 | DMT400035542 | CAACGAAGATGATCCATCTTTTCTGTTGTATTGTTTGTAATTTAGGGAGTGTAATCGCAT |
All Microarray Probes Designed to Gene DMG400013663
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44206_PI426222305 | JHI_St_60k_v1 | DMT400035541 | ATTCATGGGATATTGAGGATGAGAATGCTGAGCAGCTTATTATTGAGGAAGATAGCCGCC |
| CUST_44204_PI426222305 | JHI_St_60k_v1 | DMT400035542 | CAACGAAGATGATCCATCTTTTCTGTTGTATTGTTTGTAATTTAGGGAGTGTAATCGCAT |
Microarray Signals from CUST_44204_PI426222305
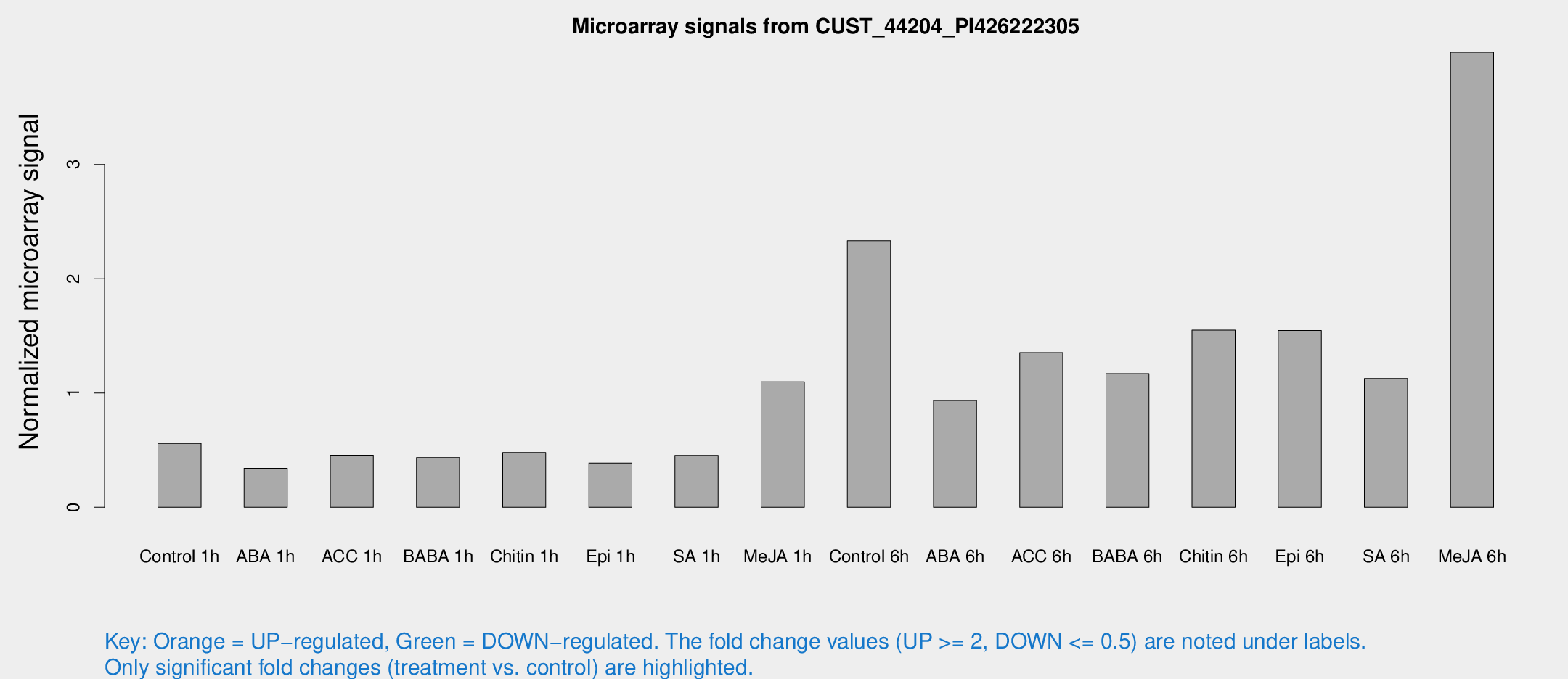
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 312.225 | 56.1117 | 0.559778 | 0.111437 |
| ABA 1h | 170.245 | 30.6393 | 0.342427 | 0.0779067 |
| ACC 1h | 455.928 | 290.774 | 0.455597 | 0.589676 |
| BABA 1h | 233.093 | 36.2068 | 0.435352 | 0.11618 |
| Chitin 1h | 271.152 | 110.213 | 0.479698 | 0.20282 |
| Epi 1h | 185.147 | 23.7104 | 0.388108 | 0.0529938 |
| SA 1h | 325.596 | 170.092 | 0.454111 | 0.266686 |
| Me-JA 1h | 490.243 | 47.6241 | 1.09833 | 0.197942 |
| Control 6h | 1445.99 | 438.081 | 2.33266 | 0.753412 |
| ABA 6h | 545.96 | 63.3418 | 0.935311 | 0.0644906 |
| ACC 6h | 859.766 | 124.129 | 1.35387 | 0.115243 |
| BABA 6h | 732.51 | 138.43 | 1.1695 | 0.217718 |
| Chitin 6h | 904.935 | 98.4785 | 1.55045 | 0.13743 |
| Epi 6h | 982.899 | 191.022 | 1.54728 | 0.457651 |
| SA 6h | 627.277 | 119.232 | 1.12672 | 0.116165 |
| Me-JA 6h | 2236.69 | 471.285 | 3.98185 | 0.509876 |
Source Transcript PGSC0003DMT400035542 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G76490.1 | +2 | 0.0 | 669 | 349/436 (80%) | hydroxy methylglutaryl CoA reductase 1 | chr1:28695801-28698206 FORWARD LENGTH=642 |
