Probe CUST_44206_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44206_PI426222305 | JHI_St_60k_v1 | DMT400035541 | ATTCATGGGATATTGAGGATGAGAATGCTGAGCAGCTTATTATTGAGGAAGATAGCCGCC |
All Microarray Probes Designed to Gene DMG400013663
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44206_PI426222305 | JHI_St_60k_v1 | DMT400035541 | ATTCATGGGATATTGAGGATGAGAATGCTGAGCAGCTTATTATTGAGGAAGATAGCCGCC |
| CUST_44204_PI426222305 | JHI_St_60k_v1 | DMT400035542 | CAACGAAGATGATCCATCTTTTCTGTTGTATTGTTTGTAATTTAGGGAGTGTAATCGCAT |
Microarray Signals from CUST_44206_PI426222305
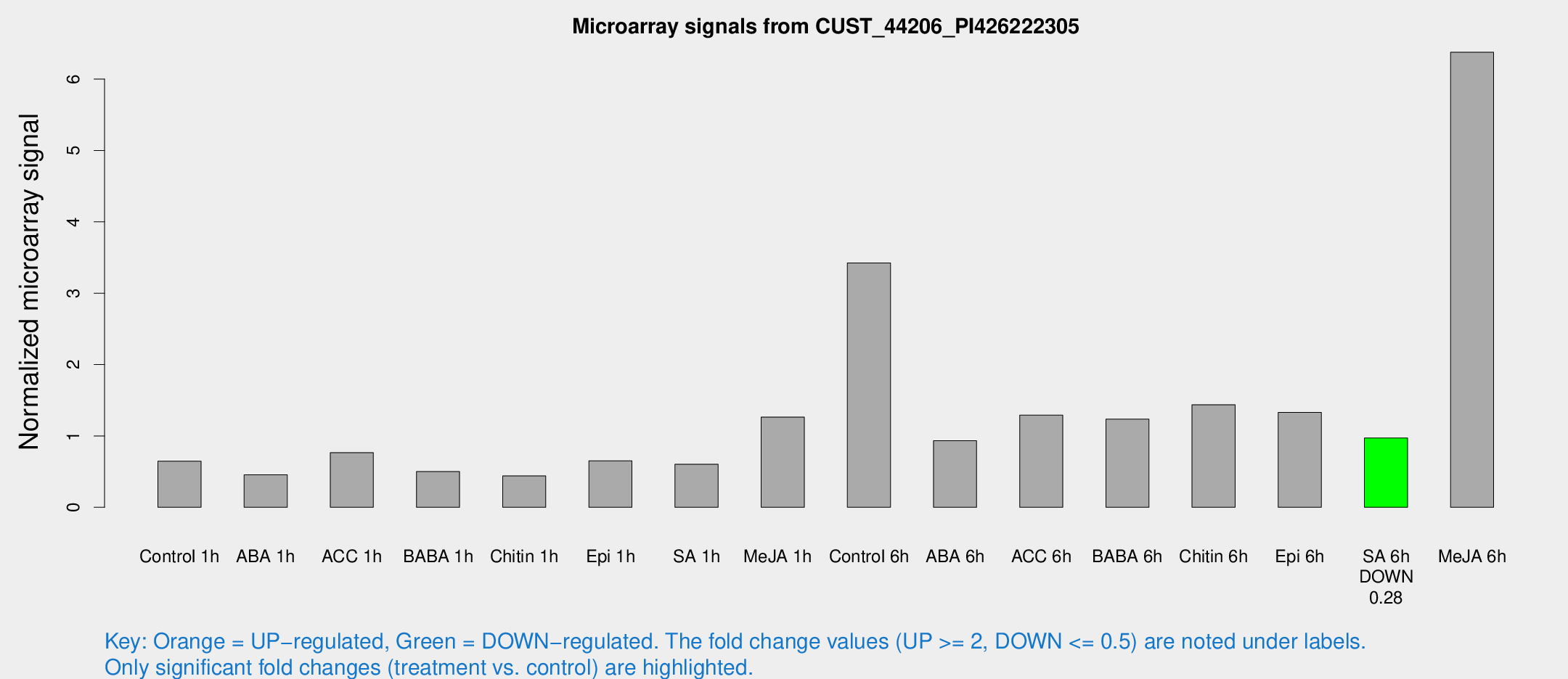
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 12.435 | 3.43837 | 0.646408 | 0.193331 |
| ABA 1h | 8.2532 | 3.24255 | 0.45503 | 0.21713 |
| ACC 1h | 18.3172 | 8.41422 | 0.766157 | 0.454212 |
| BABA 1h | 9.13182 | 3.62673 | 0.500561 | 0.214223 |
| Chitin 1h | 7.74004 | 3.36905 | 0.439584 | 0.212534 |
| Epi 1h | 13.5405 | 7.02352 | 0.65162 | 0.378473 |
| SA 1h | 13.0307 | 4.65206 | 0.603241 | 0.268951 |
| Me-JA 1h | 19.5335 | 3.62111 | 1.26473 | 0.329829 |
| Control 6h | 71.0804 | 20.4843 | 3.42449 | 0.993449 |
| ABA 6h | 23.3498 | 8.69627 | 0.933097 | 0.59424 |
| ACC 6h | 27.5211 | 4.38165 | 1.29154 | 0.265233 |
| BABA 6h | 26.883 | 5.87148 | 1.23482 | 0.335802 |
| Chitin 6h | 28.4324 | 4.1572 | 1.43686 | 0.214704 |
| Epi 6h | 33.0856 | 14.2159 | 1.32912 | 0.682626 |
| SA 6h | 18.0912 | 3.81986 | 0.970858 | 0.215059 |
| Me-JA 6h | 122.264 | 24.9268 | 6.3769 | 0.801227 |
Source Transcript PGSC0003DMT400035541 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G76490.1 | +1 | 1e-41 | 150 | 85/183 (46%) | hydroxy methylglutaryl CoA reductase 1 | chr1:28695801-28698206 FORWARD LENGTH=642 |
