Probe CUST_44118_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44118_PI426222305 | JHI_St_60k_v1 | DMT400055198 | GTTCCAGAACAGAAGAGCCAGGTAGAACTAATCACTAATGATAATCTTCACTATTGTTTT |
All Microarray Probes Designed to Gene DMG400021423
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44118_PI426222305 | JHI_St_60k_v1 | DMT400055198 | GTTCCAGAACAGAAGAGCCAGGTAGAACTAATCACTAATGATAATCTTCACTATTGTTTT |
| CUST_44129_PI426222305 | JHI_St_60k_v1 | DMT400055199 | AGAAGAGCCAGAACAAAACTGAAGCAAACAGAGGTAGACTGTGAATTCCTGAAAAAATGC |
Microarray Signals from CUST_44118_PI426222305
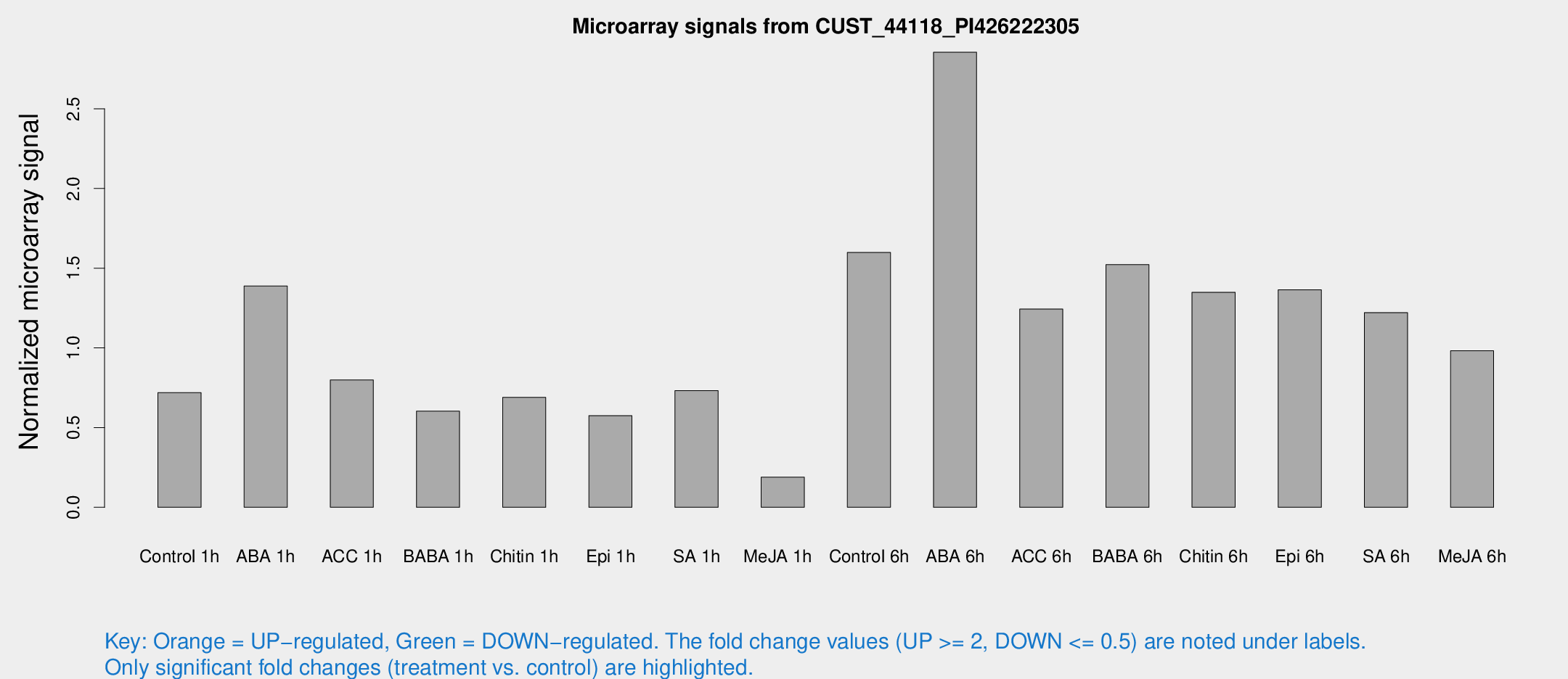
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 63.3717 | 5.3568 | 0.719815 | 0.0585434 |
| ABA 1h | 112.275 | 21.7654 | 1.38797 | 0.232763 |
| ACC 1h | 75.1763 | 15.6963 | 0.799003 | 0.127576 |
| BABA 1h | 54.7915 | 13.2482 | 0.602843 | 0.114771 |
| Chitin 1h | 54.6853 | 4.93216 | 0.689838 | 0.0602898 |
| Epi 1h | 43.8888 | 4.22081 | 0.574953 | 0.0701679 |
| SA 1h | 68.4523 | 12.7182 | 0.731826 | 0.121023 |
| Me-JA 1h | 16.8711 | 7.73671 | 0.18918 | 0.083276 |
| Control 6h | 180.826 | 75.4581 | 1.5986 | 0.893284 |
| ABA 6h | 281.763 | 63.4586 | 2.85502 | 0.600998 |
| ACC 6h | 127.367 | 18.6217 | 1.2437 | 0.084082 |
| BABA 6h | 156.971 | 32.4875 | 1.52226 | 0.315838 |
| Chitin 6h | 128.302 | 17.8777 | 1.34907 | 0.210786 |
| Epi 6h | 142.03 | 34.7682 | 1.36453 | 0.316617 |
| SA 6h | 110.856 | 25.0169 | 1.22152 | 0.182746 |
| Me-JA 6h | 100.402 | 33.577 | 0.981847 | 0.383419 |
Source Transcript PGSC0003DMT400055198 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37790.1 | +2 | 2e-63 | 133 | 73/102 (72%) | Homeobox-leucine zipper protein family | chr4:17768241-17769272 FORWARD LENGTH=278 |
