Probe CUST_44129_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44129_PI426222305 | JHI_St_60k_v1 | DMT400055199 | AGAAGAGCCAGAACAAAACTGAAGCAAACAGAGGTAGACTGTGAATTCCTGAAAAAATGC |
All Microarray Probes Designed to Gene DMG400021423
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44118_PI426222305 | JHI_St_60k_v1 | DMT400055198 | GTTCCAGAACAGAAGAGCCAGGTAGAACTAATCACTAATGATAATCTTCACTATTGTTTT |
| CUST_44129_PI426222305 | JHI_St_60k_v1 | DMT400055199 | AGAAGAGCCAGAACAAAACTGAAGCAAACAGAGGTAGACTGTGAATTCCTGAAAAAATGC |
Microarray Signals from CUST_44129_PI426222305
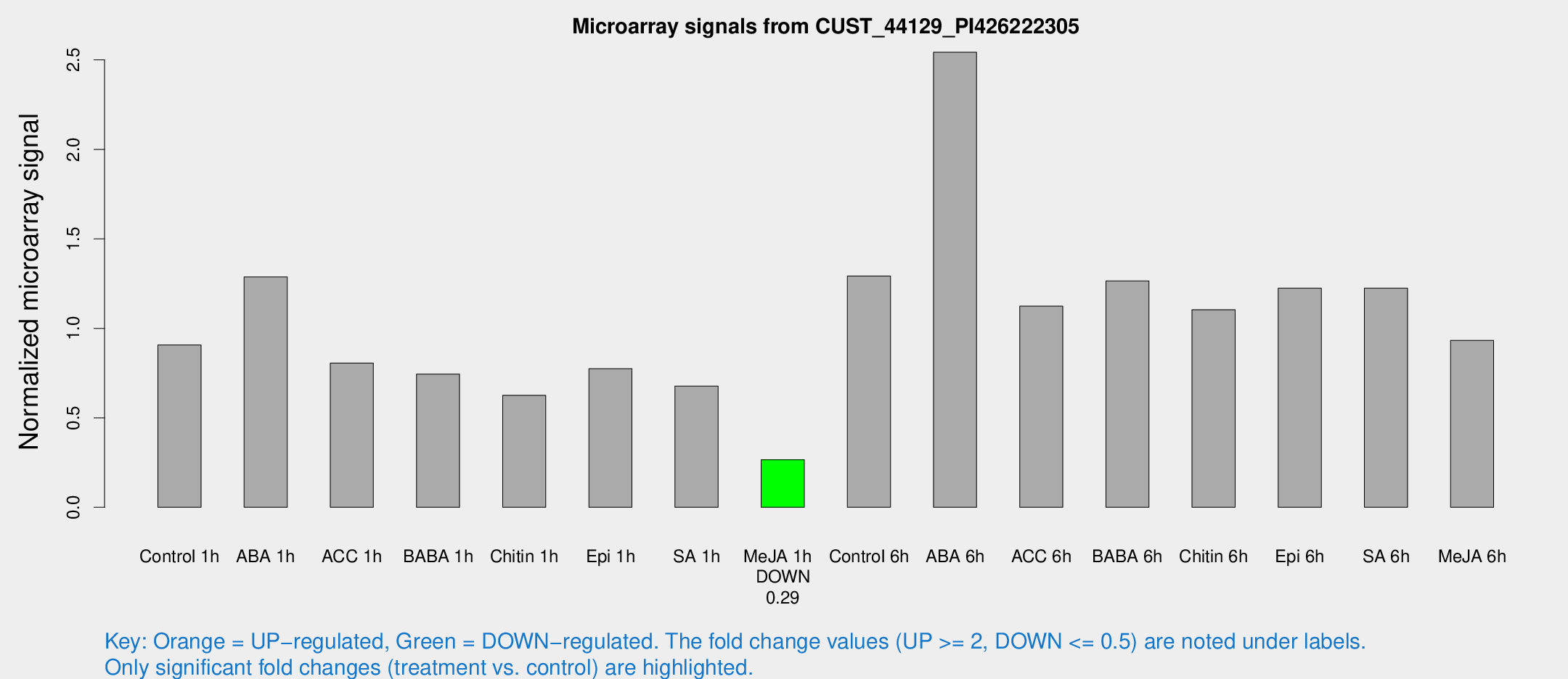
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1640.44 | 212.491 | 0.907747 | 0.0946008 |
| ABA 1h | 2076.96 | 343.045 | 1.28729 | 0.117944 |
| ACC 1h | 1604.87 | 412.355 | 0.806194 | 0.198115 |
| BABA 1h | 1331.57 | 254.059 | 0.744566 | 0.0827979 |
| Chitin 1h | 1053.49 | 234.162 | 0.625989 | 0.13215 |
| Epi 1h | 1240.46 | 231.2 | 0.775105 | 0.146567 |
| SA 1h | 1325.48 | 359.782 | 0.677272 | 0.138856 |
| Me-JA 1h | 389.33 | 35.8741 | 0.26609 | 0.0154975 |
| Control 6h | 2620.37 | 871.354 | 1.29231 | 0.408575 |
| ABA 6h | 4897.49 | 644.487 | 2.54307 | 0.230549 |
| ACC 6h | 2362.92 | 389.909 | 1.12385 | 0.127841 |
| BABA 6h | 2530.38 | 179.851 | 1.26495 | 0.0730516 |
| Chitin 6h | 2092.6 | 121.074 | 1.10403 | 0.0637652 |
| Epi 6h | 2551.25 | 476.093 | 1.22496 | 0.189884 |
| SA 6h | 2218.02 | 367.455 | 1.22463 | 0.0878714 |
| Me-JA 6h | 1704.88 | 295.647 | 0.933158 | 0.0887654 |
Source Transcript PGSC0003DMT400055199 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37790.1 | +1 | 2e-87 | 270 | 173/287 (60%) | Homeobox-leucine zipper protein family | chr4:17768241-17769272 FORWARD LENGTH=278 |
