Probe CUST_44044_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44044_PI426222305 | JHI_St_60k_v1 | DMT400065823 | TTTCAAGTACTGATGCATACGTGCGCTACATCTTCTCGTGATGTAGGTTCAGGTTCTCAG |
All Microarray Probes Designed to Gene DMG400025621
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44044_PI426222305 | JHI_St_60k_v1 | DMT400065823 | TTTCAAGTACTGATGCATACGTGCGCTACATCTTCTCGTGATGTAGGTTCAGGTTCTCAG |
| CUST_44043_PI426222305 | JHI_St_60k_v1 | DMT400065825 | GCACAACAACTTCATTGCTATCTTATGGGTTTAGGTTTTCCCCTAATTGTTTTGGTTAGC |
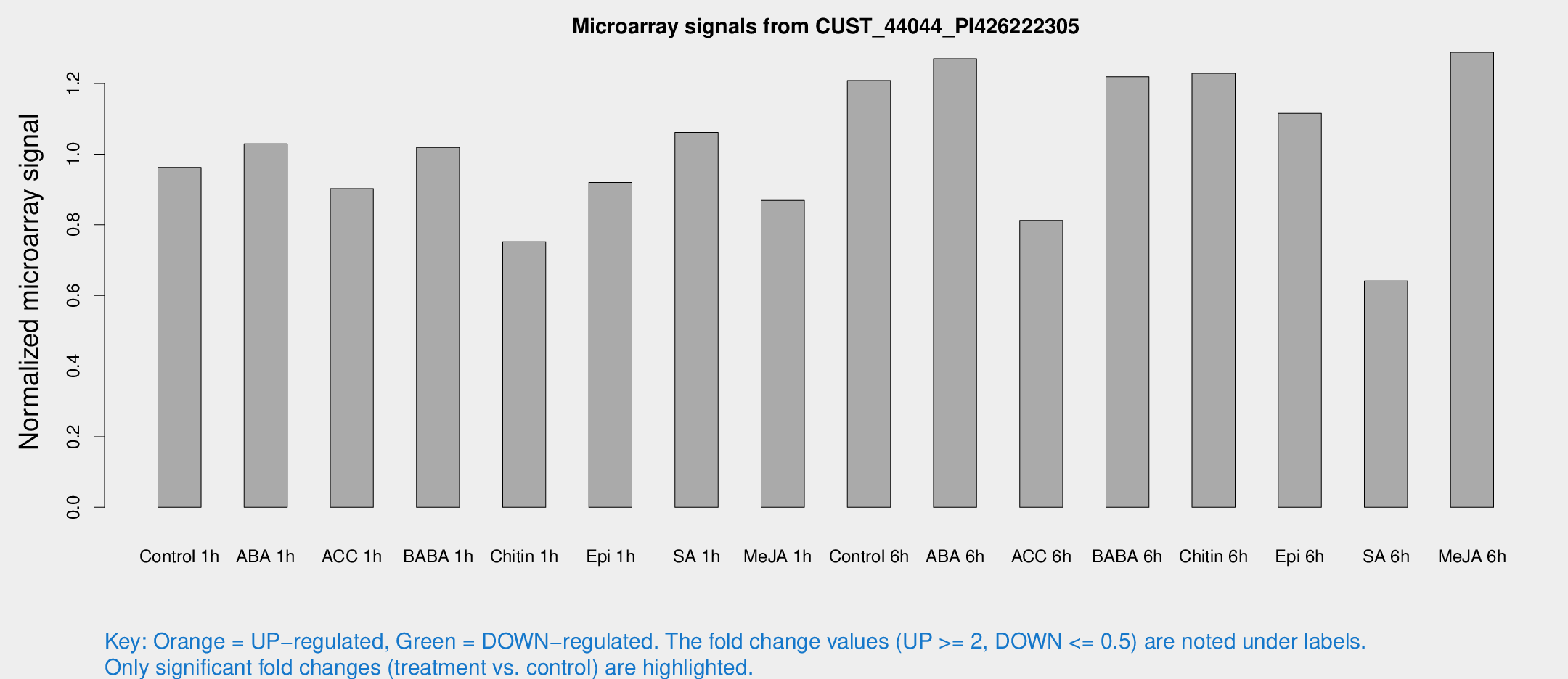
Microarray Signals from CUST_44044_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 145.249 | 8.93315 | 0.962112 | 0.0939667 |
| ABA 1h | 139.126 | 17.6413 | 1.02922 | 0.120916 |
| ACC 1h | 141.894 | 18.4467 | 0.902033 | 0.0620586 |
| BABA 1h | 153.137 | 27.2331 | 1.01893 | 0.0973593 |
| Chitin 1h | 102.178 | 6.95766 | 0.752021 | 0.0487344 |
| Epi 1h | 120.753 | 10.9707 | 0.919659 | 0.0792392 |
| SA 1h | 164.395 | 9.97995 | 1.06113 | 0.0688158 |
| Me-JA 1h | 110.049 | 18.529 | 0.868928 | 0.155488 |
| Control 6h | 193.724 | 44.3377 | 1.20831 | 0.200457 |
| ABA 6h | 211.572 | 40.6291 | 1.26972 | 0.231914 |
| ACC 6h | 141.409 | 13.0011 | 0.812337 | 0.124097 |
| BABA 6h | 211.493 | 34.2485 | 1.21905 | 0.173265 |
| Chitin 6h | 197.415 | 12.0149 | 1.22883 | 0.0740966 |
| Epi 6h | 197.346 | 42.1502 | 1.11515 | 0.262951 |
| SA 6h | 108.517 | 39.4805 | 0.640649 | 0.25204 |
| Me-JA 6h | 201.43 | 38.181 | 1.2883 | 0.186558 |
Source Transcript PGSC0003DMT400065823 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20600.1 | +1 | 3e-09 | 55 | 41/109 (38%) | Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family | chr3:7194877-7195536 FORWARD LENGTH=219 |
