Probe CUST_44043_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44043_PI426222305 | JHI_St_60k_v1 | DMT400065825 | GCACAACAACTTCATTGCTATCTTATGGGTTTAGGTTTTCCCCTAATTGTTTTGGTTAGC |
All Microarray Probes Designed to Gene DMG400025621
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44044_PI426222305 | JHI_St_60k_v1 | DMT400065823 | TTTCAAGTACTGATGCATACGTGCGCTACATCTTCTCGTGATGTAGGTTCAGGTTCTCAG |
| CUST_44043_PI426222305 | JHI_St_60k_v1 | DMT400065825 | GCACAACAACTTCATTGCTATCTTATGGGTTTAGGTTTTCCCCTAATTGTTTTGGTTAGC |
Microarray Signals from CUST_44043_PI426222305
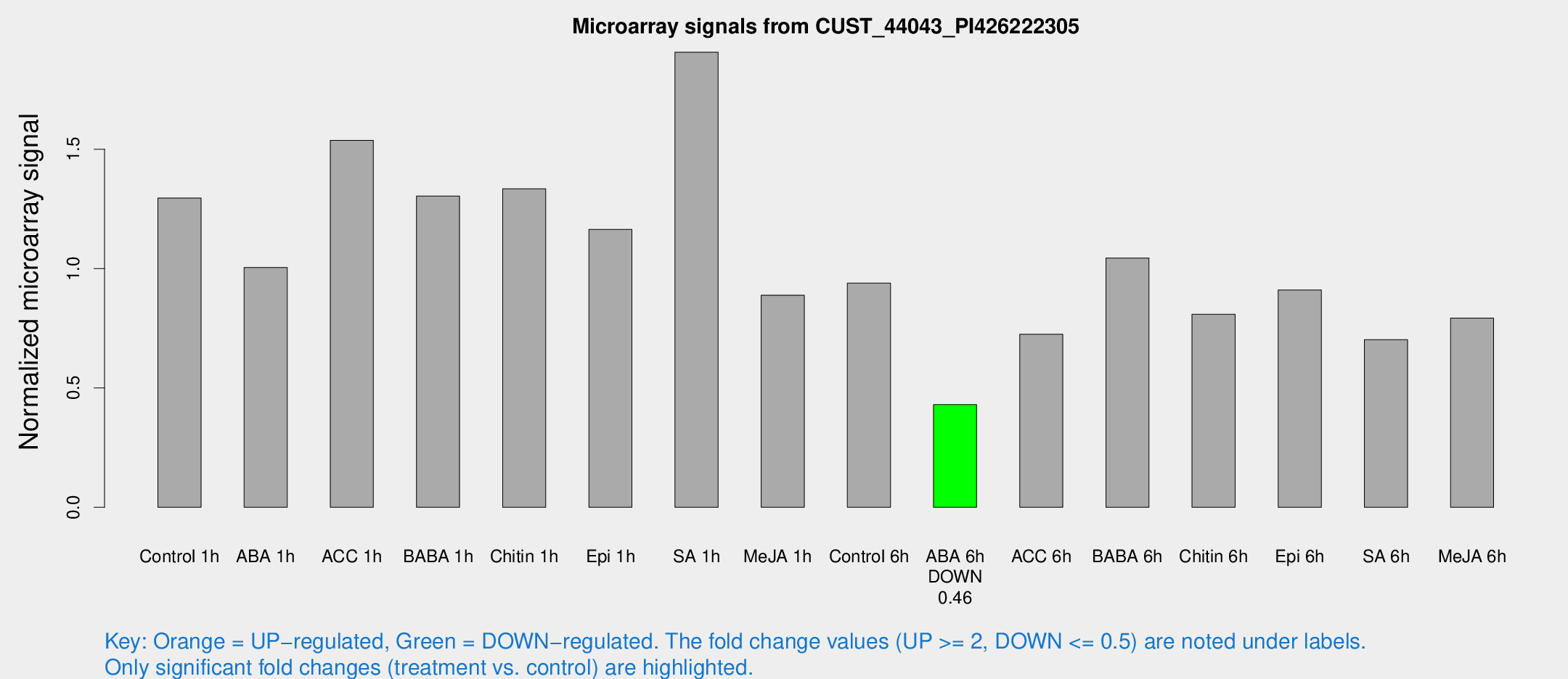
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1893.93 | 326.722 | 1.29499 | 0.132528 |
| ABA 1h | 1264.03 | 73.1321 | 1.00404 | 0.0606808 |
| ACC 1h | 2449.33 | 628.785 | 1.53697 | 0.379598 |
| BABA 1h | 1830.1 | 282.062 | 1.30379 | 0.108778 |
| Chitin 1h | 1728.08 | 193.856 | 1.33443 | 0.133312 |
| Epi 1h | 1434.26 | 82.9816 | 1.16408 | 0.0672737 |
| SA 1h | 2867.83 | 461.907 | 1.90601 | 0.233026 |
| Me-JA 1h | 1038.31 | 82.3685 | 0.888476 | 0.0514001 |
| Control 6h | 1431.3 | 344.831 | 0.939063 | 0.16686 |
| ABA 6h | 658.114 | 74.196 | 0.430106 | 0.0249781 |
| ACC 6h | 1192.93 | 115.551 | 0.724777 | 0.0426111 |
| BABA 6h | 1672.45 | 134.054 | 1.04411 | 0.0603423 |
| Chitin 6h | 1225.1 | 70.9349 | 0.808486 | 0.0467621 |
| Epi 6h | 1469.88 | 124.098 | 0.910078 | 0.132481 |
| SA 6h | 1025.23 | 189.249 | 0.702444 | 0.0679107 |
| Me-JA 6h | 1185.8 | 285.638 | 0.792281 | 0.123274 |
Source Transcript PGSC0003DMT400065825 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20600.1 | +2 | 1e-32 | 122 | 76/181 (42%) | Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family | chr3:7194877-7195536 FORWARD LENGTH=219 |
