Probe CUST_44037_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44037_PI426222305 | JHI_St_60k_v1 | DMT400016515 | CCTGTGGTTTGATATTGCACTAGGACCAAAAATGTATCCTTACAAAGTTGTATACAGAAA |
All Microarray Probes Designed to Gene DMG400006450
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44014_PI426222305 | JHI_St_60k_v1 | DMT400016514 | CCTGTGGTTTGATATTGCACTAGGACCAAAAATGTATCCTTACAAAGTTGTATACAGAAA |
| CUST_44037_PI426222305 | JHI_St_60k_v1 | DMT400016515 | CCTGTGGTTTGATATTGCACTAGGACCAAAAATGTATCCTTACAAAGTTGTATACAGAAA |
Microarray Signals from CUST_44037_PI426222305
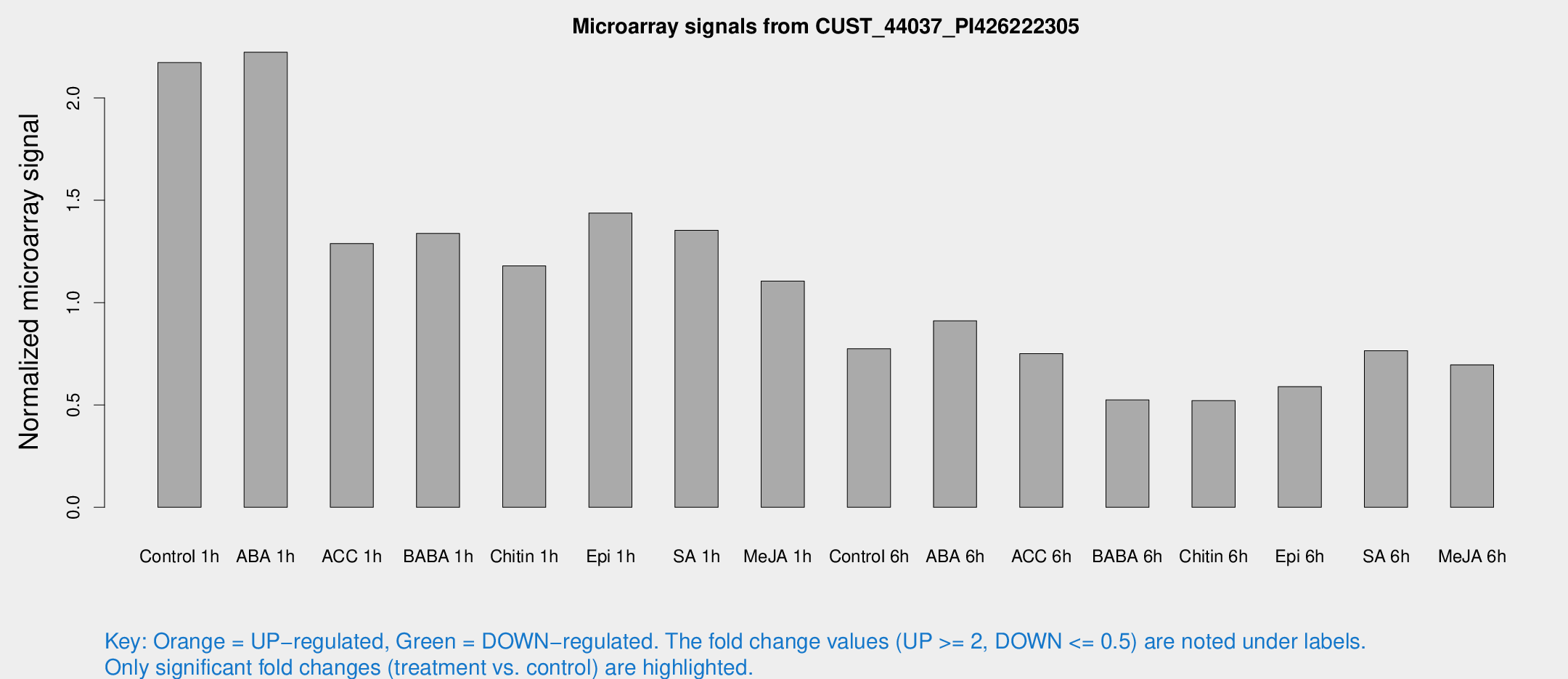
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3518.66 | 562.234 | 2.17246 | 0.205858 |
| ABA 1h | 3163.79 | 421.626 | 2.22328 | 0.248253 |
| ACC 1h | 2262.67 | 603.702 | 1.28869 | 0.300259 |
| BABA 1h | 2215.05 | 567.124 | 1.3378 | 0.27988 |
| Chitin 1h | 1707.46 | 243.994 | 1.17918 | 0.0735345 |
| Epi 1h | 1981.62 | 183.022 | 1.43752 | 0.128143 |
| SA 1h | 2248.81 | 329.895 | 1.35332 | 0.147649 |
| Me-JA 1h | 1451.08 | 196.124 | 1.10531 | 0.0638692 |
| Control 6h | 1286.46 | 280.306 | 0.774982 | 0.125388 |
| ABA 6h | 1571.05 | 252.179 | 0.911161 | 0.102893 |
| ACC 6h | 1388.5 | 190.854 | 0.750679 | 0.102446 |
| BABA 6h | 934.724 | 82.0058 | 0.524785 | 0.0330986 |
| Chitin 6h | 879.874 | 64.9717 | 0.52086 | 0.0301605 |
| Epi 6h | 1061.62 | 115.284 | 0.589186 | 0.0461417 |
| SA 6h | 1265.35 | 289.674 | 0.764476 | 0.108628 |
| Me-JA 6h | 1105.71 | 112.605 | 0.696063 | 0.0402488 |
Source Transcript PGSC0003DMT400016515 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G43850.1 | +2 | 3e-112 | 327 | 146/187 (78%) | RmlC-like cupins superfamily protein | chr5:17627364-17629122 REVERSE LENGTH=187 |
