Probe CUST_44014_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44014_PI426222305 | JHI_St_60k_v1 | DMT400016514 | CCTGTGGTTTGATATTGCACTAGGACCAAAAATGTATCCTTACAAAGTTGTATACAGAAA |
All Microarray Probes Designed to Gene DMG400006450
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44014_PI426222305 | JHI_St_60k_v1 | DMT400016514 | CCTGTGGTTTGATATTGCACTAGGACCAAAAATGTATCCTTACAAAGTTGTATACAGAAA |
| CUST_44037_PI426222305 | JHI_St_60k_v1 | DMT400016515 | CCTGTGGTTTGATATTGCACTAGGACCAAAAATGTATCCTTACAAAGTTGTATACAGAAA |
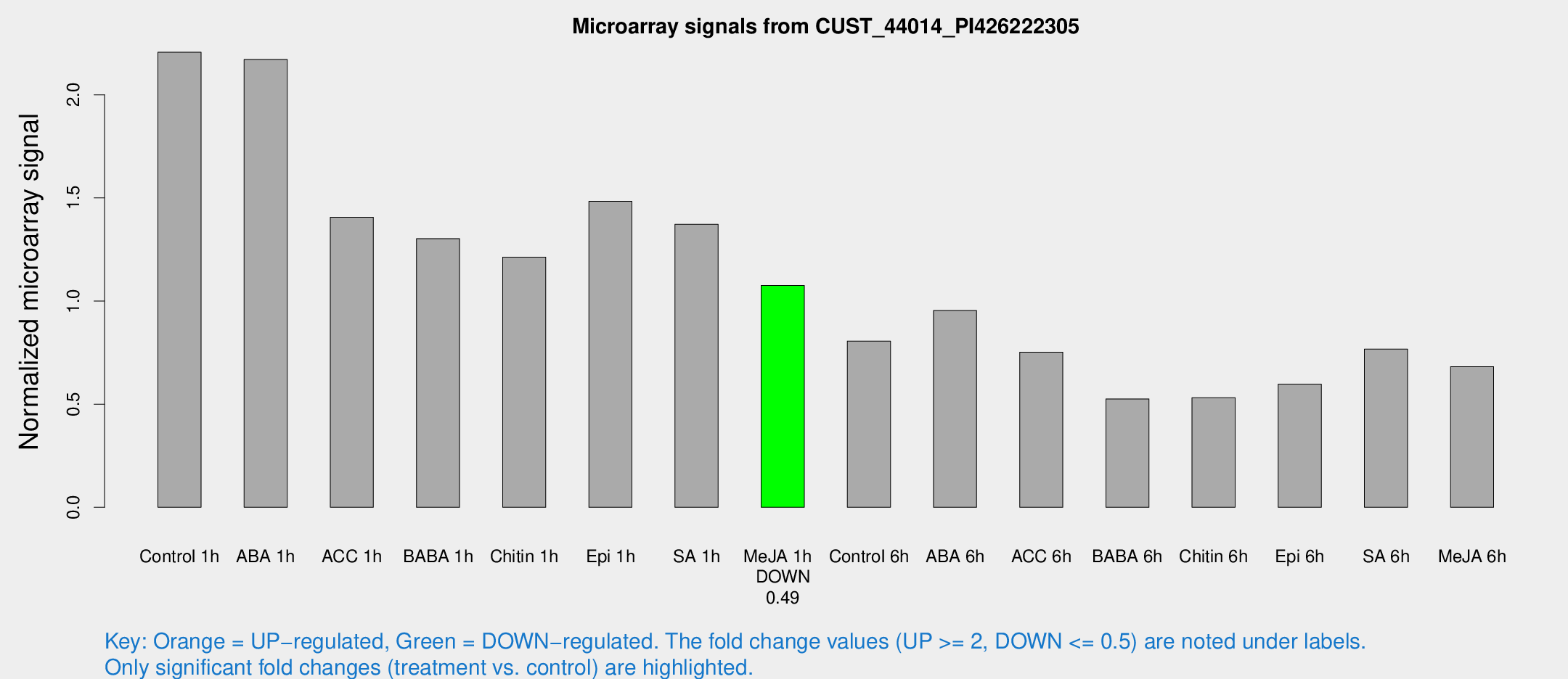
Microarray Signals from CUST_44014_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3364.89 | 354.076 | 2.2062 | 0.127399 |
| ABA 1h | 2955.97 | 405.109 | 2.17135 | 0.258647 |
| ACC 1h | 2313.71 | 530.874 | 1.40606 | 0.264601 |
| BABA 1h | 2035.32 | 480.728 | 1.30261 | 0.235379 |
| Chitin 1h | 1668.02 | 195.783 | 1.21308 | 0.0700876 |
| Epi 1h | 1950.79 | 166.974 | 1.48308 | 0.134934 |
| SA 1h | 2185.54 | 341.135 | 1.37158 | 0.156838 |
| Me-JA 1h | 1355.88 | 201.135 | 1.07588 | 0.0702134 |
| Control 6h | 1280.53 | 287.794 | 0.806069 | 0.138665 |
| ABA 6h | 1560.66 | 213.94 | 0.954098 | 0.0817456 |
| ACC 6h | 1318.97 | 142.259 | 0.752275 | 0.0861214 |
| BABA 6h | 891.86 | 62.1929 | 0.525405 | 0.030443 |
| Chitin 6h | 857.514 | 53.2986 | 0.53171 | 0.0312755 |
| Epi 6h | 1031.2 | 120.715 | 0.596964 | 0.039396 |
| SA 6h | 1214.69 | 276.65 | 0.766639 | 0.109255 |
| Me-JA 6h | 1031.6 | 85.1323 | 0.681952 | 0.039458 |
Source Transcript PGSC0003DMT400016514 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G43850.1 | +3 | 7e-110 | 320 | 143/183 (78%) | RmlC-like cupins superfamily protein | chr5:17627364-17629122 REVERSE LENGTH=187 |
