Probe CUST_43925_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43925_PI426222305 | JHI_St_60k_v1 | DMT400045360 | CAAGACTAGGCGATGACAGTGTAGCCATTGCTACTGTTAATTAGAAATATGTATAAAGAT |
All Microarray Probes Designed to Gene DMG400017602
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43925_PI426222305 | JHI_St_60k_v1 | DMT400045360 | CAAGACTAGGCGATGACAGTGTAGCCATTGCTACTGTTAATTAGAAATATGTATAAAGAT |
| CUST_43924_PI426222305 | JHI_St_60k_v1 | DMT400045361 | CAAGACTAGGCGATGACAGTGTAGCCATTGCTACTGTTAATTAGAAATATGTATAAAGAT |
Microarray Signals from CUST_43925_PI426222305
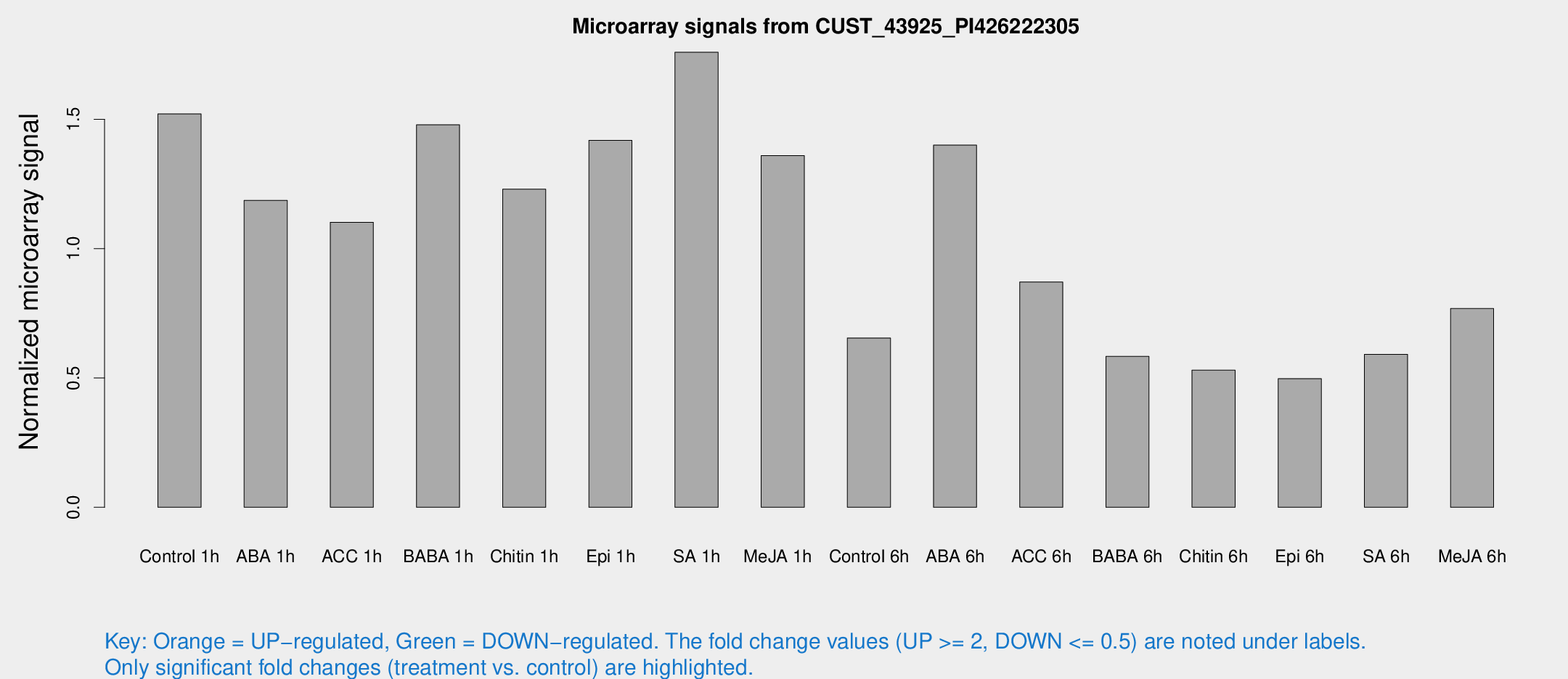
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 103.269 | 26.0149 | 1.52078 | 0.314245 |
| ABA 1h | 71.6091 | 18.9859 | 1.18672 | 0.221127 |
| ACC 1h | 81.5109 | 23.9839 | 1.10201 | 0.341958 |
| BABA 1h | 101.693 | 31.4312 | 1.47887 | 0.3816 |
| Chitin 1h | 71.2688 | 6.28889 | 1.22986 | 0.0947689 |
| Epi 1h | 78.6324 | 5.60925 | 1.41847 | 0.101039 |
| SA 1h | 118.682 | 18.9311 | 1.75929 | 0.181242 |
| Me-JA 1h | 71.7719 | 7.72868 | 1.3594 | 0.106864 |
| Control 6h | 46.6204 | 12.9683 | 0.654627 | 0.172977 |
| ABA 6h | 100.274 | 21.6466 | 1.40022 | 0.258683 |
| ACC 6h | 64.1918 | 5.90652 | 0.871068 | 0.0941375 |
| BABA 6h | 42.1015 | 4.93271 | 0.58345 | 0.0683439 |
| Chitin 6h | 37.1607 | 6.48905 | 0.530087 | 0.100078 |
| Epi 6h | 39.5806 | 10.8129 | 0.497567 | 0.21989 |
| SA 6h | 39.5303 | 9.12312 | 0.590977 | 0.168301 |
| Me-JA 6h | 50.0608 | 7.16484 | 0.768468 | 0.0752055 |
Source Transcript PGSC0003DMT400045360 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37390.1 | +1 | 7e-38 | 142 | 116/276 (42%) | Chloroplast-targeted copper chaperone protein | chr2:15694300-15695461 FORWARD LENGTH=259 |
