Probe CUST_43924_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43924_PI426222305 | JHI_St_60k_v1 | DMT400045361 | CAAGACTAGGCGATGACAGTGTAGCCATTGCTACTGTTAATTAGAAATATGTATAAAGAT |
All Microarray Probes Designed to Gene DMG400017602
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43925_PI426222305 | JHI_St_60k_v1 | DMT400045360 | CAAGACTAGGCGATGACAGTGTAGCCATTGCTACTGTTAATTAGAAATATGTATAAAGAT |
| CUST_43924_PI426222305 | JHI_St_60k_v1 | DMT400045361 | CAAGACTAGGCGATGACAGTGTAGCCATTGCTACTGTTAATTAGAAATATGTATAAAGAT |
Microarray Signals from CUST_43924_PI426222305
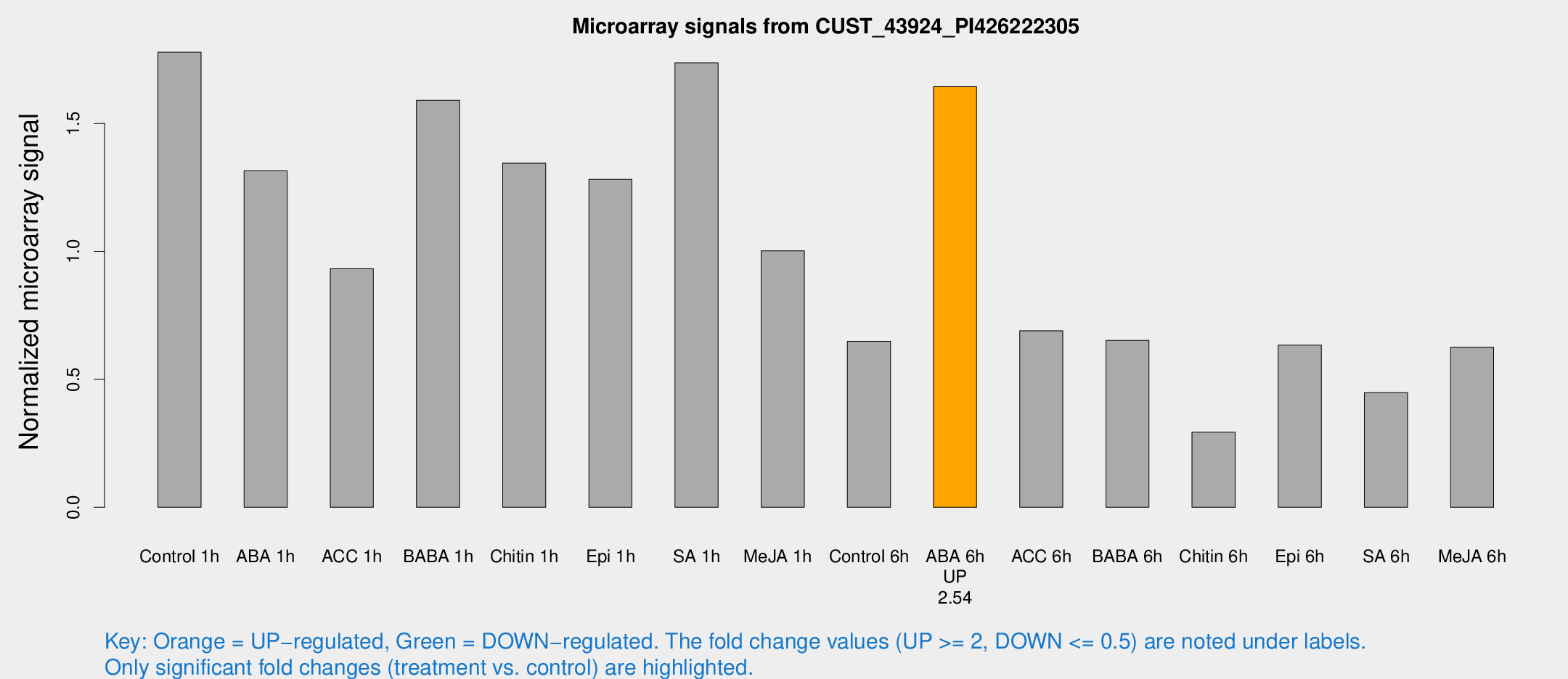
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 123.007 | 19.1481 | 1.77854 | 0.179442 |
| ABA 1h | 84.4537 | 22.6727 | 1.31524 | 0.264009 |
| ACC 1h | 82.0631 | 30.2802 | 0.93261 | 0.523539 |
| BABA 1h | 114.382 | 36.9427 | 1.59051 | 0.399181 |
| Chitin 1h | 82.8849 | 9.18069 | 1.34499 | 0.0970621 |
| Epi 1h | 75.4054 | 5.58091 | 1.28141 | 0.0949441 |
| SA 1h | 121.157 | 7.91304 | 1.73699 | 0.146642 |
| Me-JA 1h | 56.0057 | 6.08457 | 1.00233 | 0.107463 |
| Control 6h | 45.6564 | 8.54948 | 0.64844 | 0.0706684 |
| ABA 6h | 121.341 | 18.5155 | 1.64422 | 0.218526 |
| ACC 6h | 54.0224 | 5.43265 | 0.690053 | 0.0684156 |
| BABA 6h | 49.9898 | 5.42671 | 0.652081 | 0.067539 |
| Chitin 6h | 24.6668 | 10.1668 | 0.293816 | 0.113051 |
| Epi 6h | 49.8085 | 8.90054 | 0.633889 | 0.12846 |
| SA 6h | 30.2967 | 4.36899 | 0.448266 | 0.0658689 |
| Me-JA 6h | 46.0372 | 11.8708 | 0.626151 | 0.15463 |
Source Transcript PGSC0003DMT400045361 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37390.1 | +1 | 4e-16 | 80 | 82/235 (35%) | Chloroplast-targeted copper chaperone protein | chr2:15694300-15695461 FORWARD LENGTH=259 |
