Probe CUST_43282_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43282_PI426222305 | JHI_St_60k_v1 | DMT400064434 | CTGGTATTTTTTGCTCTACAACTGAAGTAATATCCCATTATGTTGCTCGGATTCTTCAAA |
All Microarray Probes Designed to Gene DMG400025034
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43282_PI426222305 | JHI_St_60k_v1 | DMT400064434 | CTGGTATTTTTTGCTCTACAACTGAAGTAATATCCCATTATGTTGCTCGGATTCTTCAAA |
| CUST_43275_PI426222305 | JHI_St_60k_v1 | DMT400064433 | TCAAGAATGGAGAGAAGAAAGACGCGGTGATTGGTGCAGTTCCAAAAACAACATTGGCCA |
Microarray Signals from CUST_43282_PI426222305
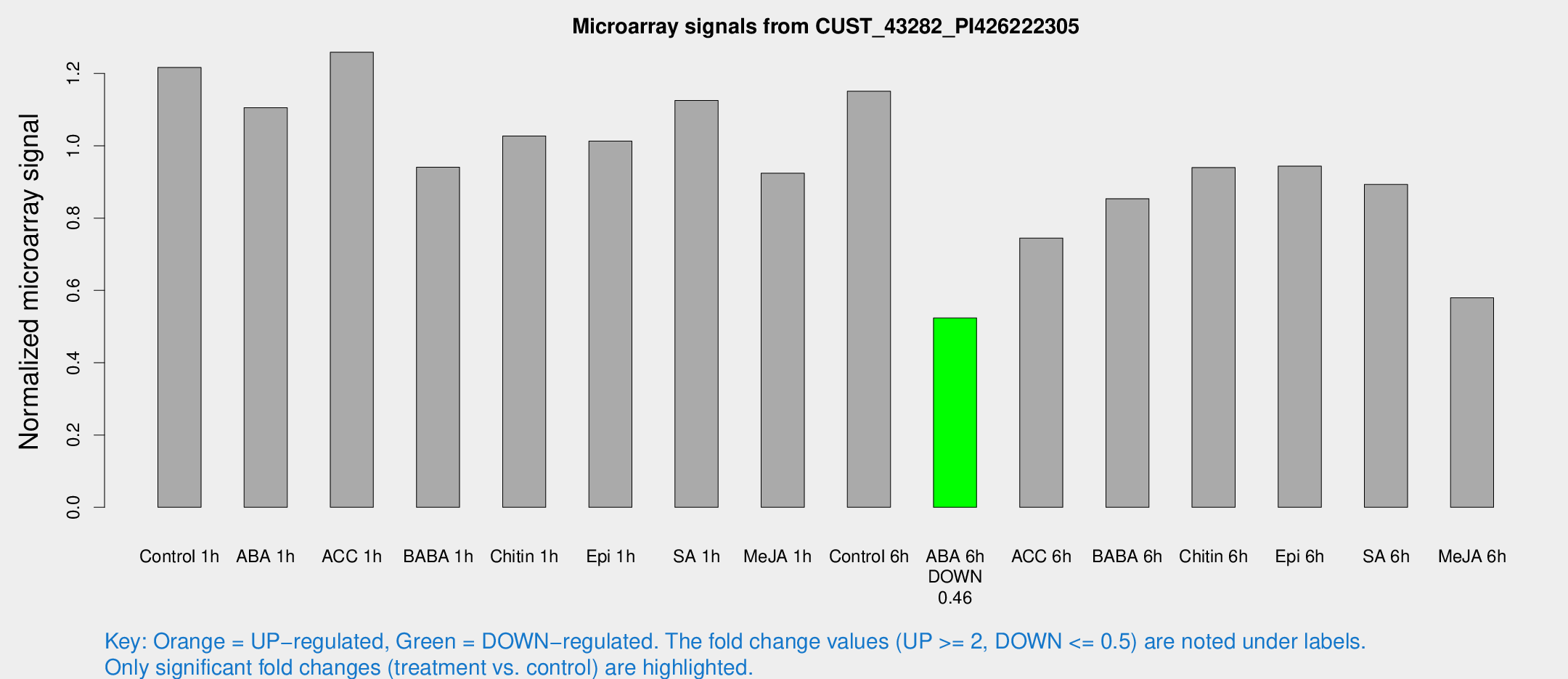
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2445.53 | 244.555 | 1.21651 | 0.0702545 |
| ABA 1h | 1959.44 | 155.084 | 1.10556 | 0.107723 |
| ACC 1h | 2574.12 | 148.719 | 1.25884 | 0.0805363 |
| BABA 1h | 1867.04 | 318.312 | 0.940666 | 0.0799222 |
| Chitin 1h | 1849.09 | 136.663 | 1.0273 | 0.0625773 |
| Epi 1h | 1751.11 | 106.79 | 1.01282 | 0.0693097 |
| SA 1h | 2314.79 | 174.308 | 1.12536 | 0.0649913 |
| Me-JA 1h | 1510.3 | 116.655 | 0.924089 | 0.0925088 |
| Control 6h | 2324.94 | 249.032 | 1.15114 | 0.0664839 |
| ABA 6h | 1114.45 | 82.8033 | 0.523863 | 0.0302902 |
| ACC 6h | 1737.19 | 259.717 | 0.744784 | 0.0430341 |
| BABA 6h | 1960.56 | 322.448 | 0.853731 | 0.18016 |
| Chitin 6h | 2008.48 | 183.995 | 0.939885 | 0.0578045 |
| Epi 6h | 2118.24 | 122.38 | 0.94369 | 0.064928 |
| SA 6h | 1829.74 | 335.919 | 0.892937 | 0.0788709 |
| Me-JA 6h | 1196.19 | 232.721 | 0.579804 | 0.068773 |
Source Transcript PGSC0003DMT400064434 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G03520.1 | +1 | 2e-48 | 163 | 85/155 (55%) | Thioredoxin superfamily protein | chr4:1562585-1564055 REVERSE LENGTH=186 |
