Probe CUST_43275_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43275_PI426222305 | JHI_St_60k_v1 | DMT400064433 | TCAAGAATGGAGAGAAGAAAGACGCGGTGATTGGTGCAGTTCCAAAAACAACATTGGCCA |
All Microarray Probes Designed to Gene DMG400025034
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43282_PI426222305 | JHI_St_60k_v1 | DMT400064434 | CTGGTATTTTTTGCTCTACAACTGAAGTAATATCCCATTATGTTGCTCGGATTCTTCAAA |
| CUST_43275_PI426222305 | JHI_St_60k_v1 | DMT400064433 | TCAAGAATGGAGAGAAGAAAGACGCGGTGATTGGTGCAGTTCCAAAAACAACATTGGCCA |
Microarray Signals from CUST_43275_PI426222305
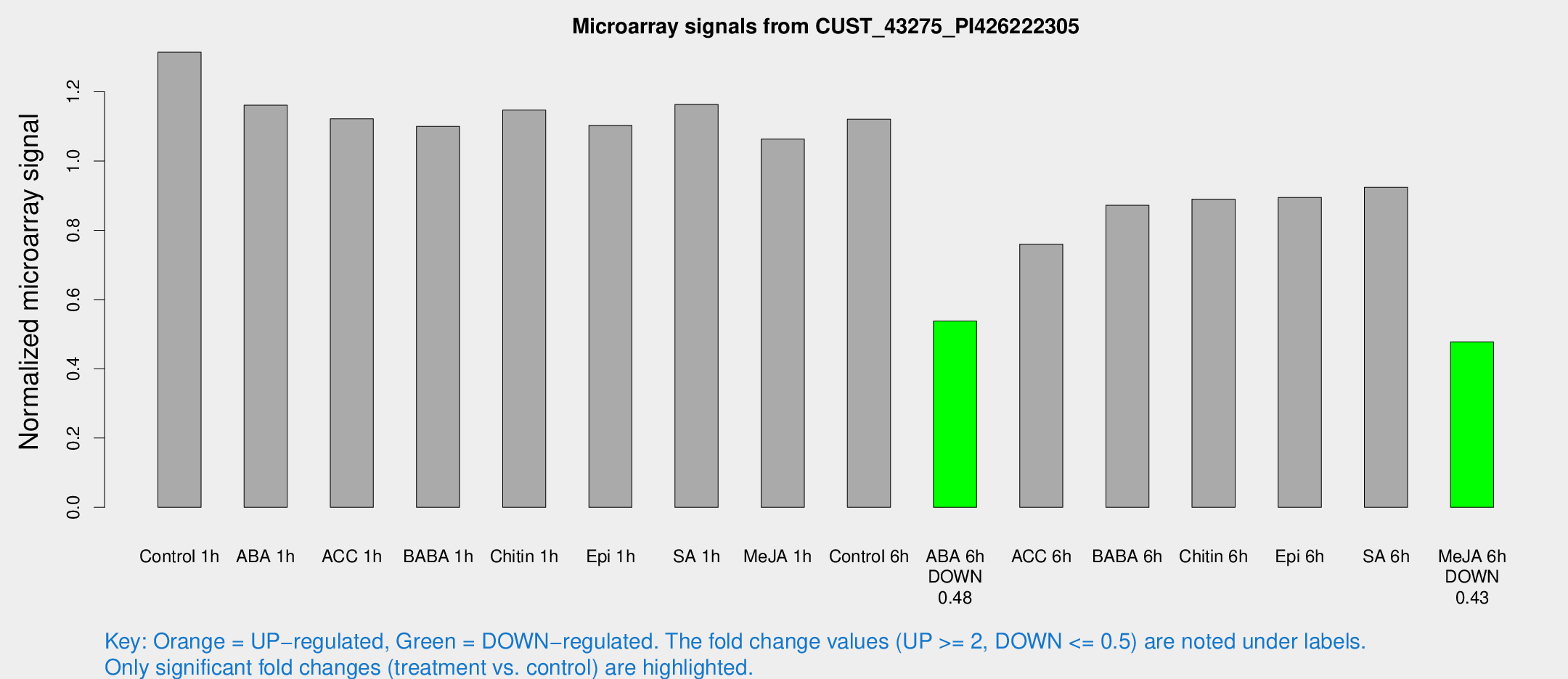
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 25123.7 | 1856.47 | 1.31424 | 0.0758777 |
| ABA 1h | 19666.8 | 1571.76 | 1.16124 | 0.126276 |
| ACC 1h | 22299 | 2822.68 | 1.12182 | 0.069248 |
| BABA 1h | 21141.2 | 4242.56 | 1.1002 | 0.135889 |
| Chitin 1h | 19850.9 | 2171.7 | 1.14709 | 0.11581 |
| Epi 1h | 18251.8 | 1313.99 | 1.10289 | 0.0696539 |
| SA 1h | 23725.3 | 4497.79 | 1.16363 | 0.198384 |
| Me-JA 1h | 16561.5 | 966.201 | 1.06313 | 0.0797444 |
| Control 6h | 21918.5 | 3448.4 | 1.12113 | 0.112822 |
| ABA 6h | 10905.5 | 631.339 | 0.538004 | 0.0310621 |
| ACC 6h | 16901.8 | 2336.77 | 0.760137 | 0.0438868 |
| BABA 6h | 18806.8 | 2043.86 | 0.87232 | 0.126959 |
| Chitin 6h | 18245.8 | 1960.71 | 0.89023 | 0.0756721 |
| Epi 6h | 19377 | 1882 | 0.894884 | 0.061205 |
| SA 6h | 17936.8 | 3027.52 | 0.923898 | 0.0663478 |
| Me-JA 6h | 9346.17 | 1737.65 | 0.47765 | 0.049335 |
Source Transcript PGSC0003DMT400064433 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G03680.1 | +1 | 8e-25 | 96 | 44/62 (71%) | thioredoxin M-type 1 | chr1:916990-917865 REVERSE LENGTH=179 |
