Probe CUST_43213_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43213_PI426222305 | JHI_St_60k_v1 | DMT400042698 | AGACAGTTGCAGTAAATCTTCATCCTCTATAGTTGAGAAGACCGTATGTCAGACCACGGC |
All Microarray Probes Designed to Gene DMG400016568
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43213_PI426222305 | JHI_St_60k_v1 | DMT400042698 | AGACAGTTGCAGTAAATCTTCATCCTCTATAGTTGAGAAGACCGTATGTCAGACCACGGC |
| CUST_43189_PI426222305 | JHI_St_60k_v1 | DMT400042699 | GGACAACATTCTTTTTGTACATGCTAGCTCAAAAACAGTTTTTAGAGAGGCTGACACTGA |
Microarray Signals from CUST_43213_PI426222305
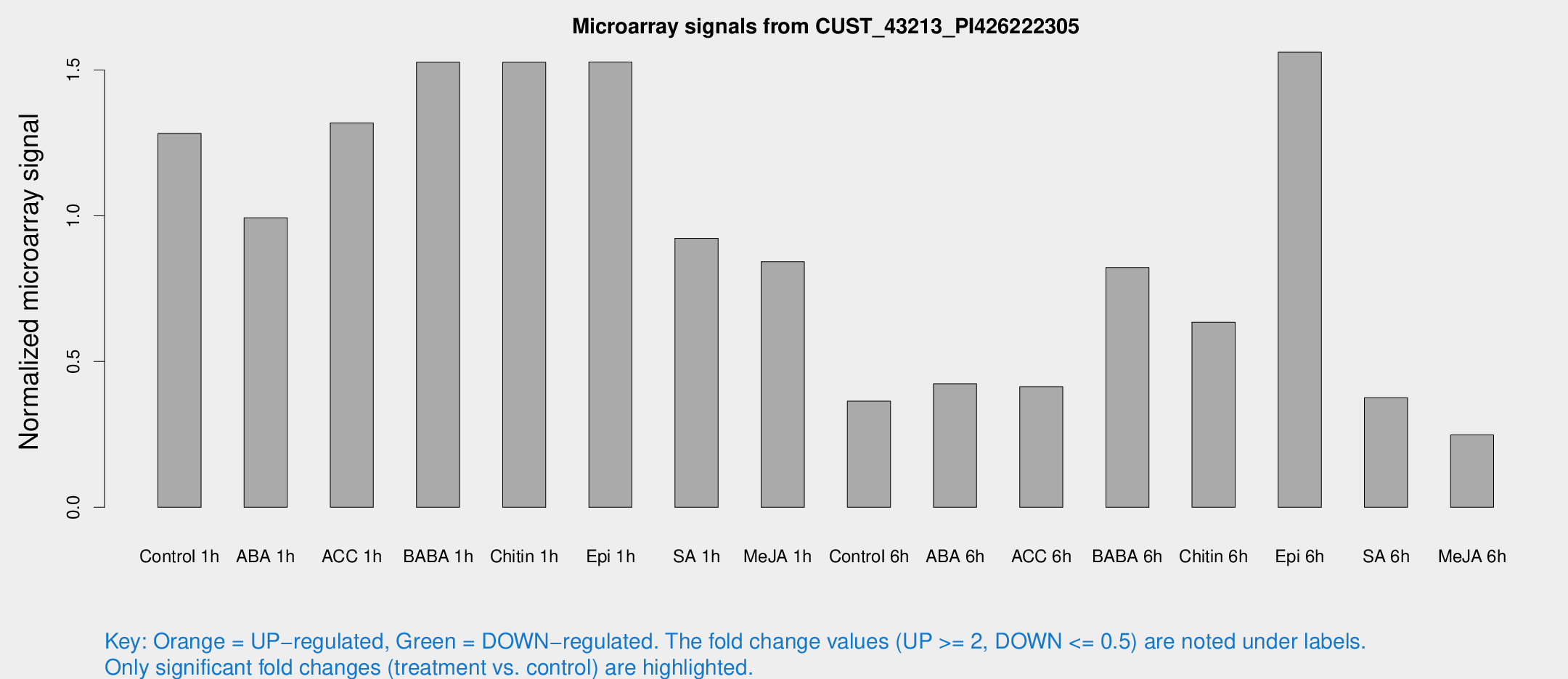
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 262.904 | 44.2137 | 1.28221 | 0.133115 |
| ABA 1h | 184.953 | 41.6736 | 0.992711 | 0.174979 |
| ACC 1h | 276.609 | 41.3346 | 1.31846 | 0.118145 |
| BABA 1h | 303.164 | 53.0534 | 1.52648 | 0.140178 |
| Chitin 1h | 287.686 | 67.8691 | 1.5265 | 0.370014 |
| Epi 1h | 272.763 | 52.2234 | 1.52716 | 0.247721 |
| SA 1h | 193.134 | 29.0351 | 0.922496 | 0.12123 |
| Me-JA 1h | 138.982 | 17.1702 | 0.842127 | 0.0529344 |
| Control 6h | 111.262 | 49.3508 | 0.363971 | 0.366769 |
| ABA 6h | 108.588 | 44.7051 | 0.423477 | 0.241822 |
| ACC 6h | 113.498 | 50.8368 | 0.413568 | 0.121096 |
| BABA 6h | 193.469 | 46.4353 | 0.822385 | 0.162006 |
| Chitin 6h | 143.78 | 36.5778 | 0.634359 | 0.163746 |
| Epi 6h | 356.754 | 46.0729 | 1.56072 | 0.292497 |
| SA 6h | 75.1387 | 9.09093 | 0.375605 | 0.0918154 |
| Me-JA 6h | 64.6222 | 31.0075 | 0.248058 | 0.1495 |
Source Transcript PGSC0003DMT400042698 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G29510.1 | +1 | 8e-07 | 45 | 20/31 (65%) | SAUR-like auxin-responsive protein family | chr1:10322683-10323114 FORWARD LENGTH=143 |
