Probe CUST_43189_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43189_PI426222305 | JHI_St_60k_v1 | DMT400042699 | GGACAACATTCTTTTTGTACATGCTAGCTCAAAAACAGTTTTTAGAGAGGCTGACACTGA |
All Microarray Probes Designed to Gene DMG400016568
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43213_PI426222305 | JHI_St_60k_v1 | DMT400042698 | AGACAGTTGCAGTAAATCTTCATCCTCTATAGTTGAGAAGACCGTATGTCAGACCACGGC |
| CUST_43189_PI426222305 | JHI_St_60k_v1 | DMT400042699 | GGACAACATTCTTTTTGTACATGCTAGCTCAAAAACAGTTTTTAGAGAGGCTGACACTGA |
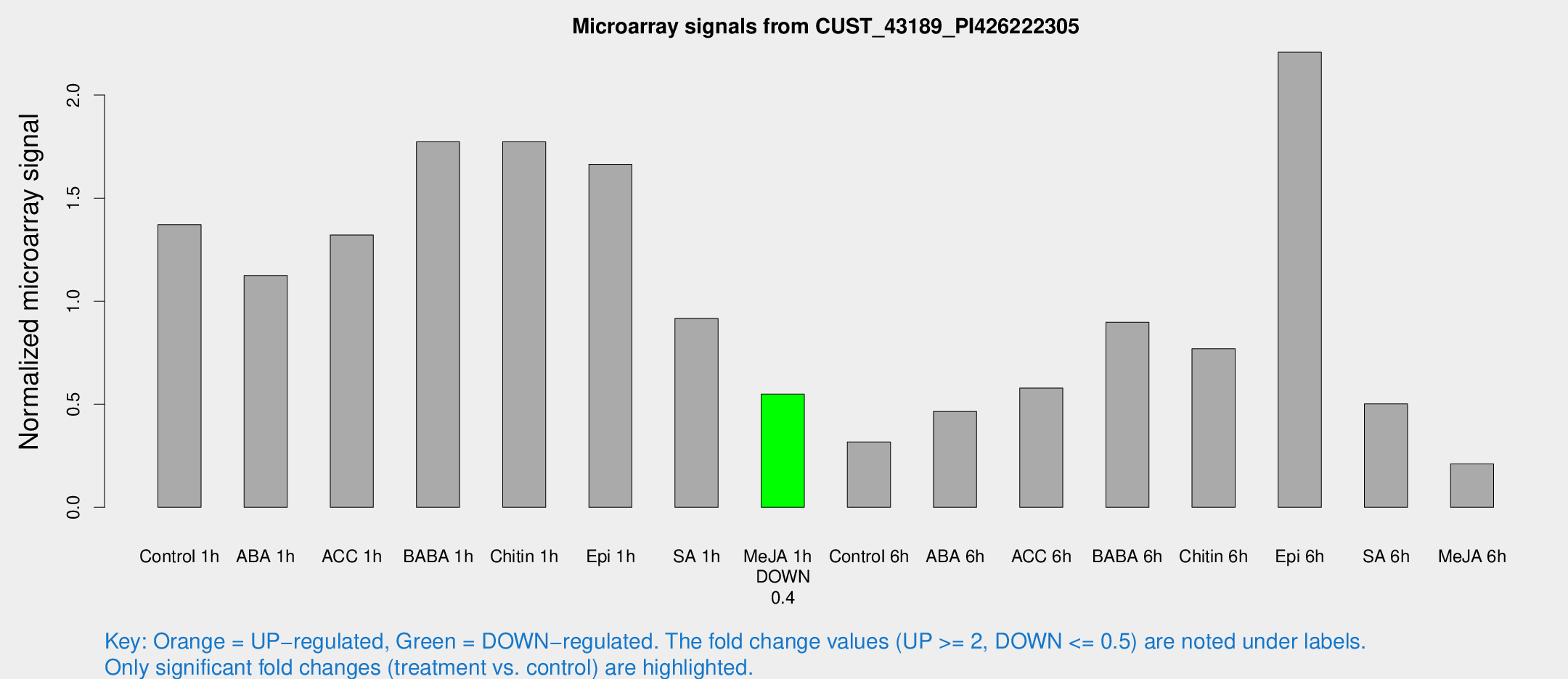
Microarray Signals from CUST_43189_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 163.39 | 17.5257 | 1.37151 | 0.084166 |
| ABA 1h | 131.487 | 41.2473 | 1.12452 | 0.339712 |
| ACC 1h | 167.366 | 32.6571 | 1.32133 | 0.195362 |
| BABA 1h | 210.285 | 40.7166 | 1.77348 | 0.201483 |
| Chitin 1h | 195.033 | 38.4752 | 1.77352 | 0.420477 |
| Epi 1h | 172.628 | 23.0457 | 1.66438 | 0.16987 |
| SA 1h | 112.85 | 14.3099 | 0.916419 | 0.120023 |
| Me-JA 1h | 53.8538 | 7.47687 | 0.549524 | 0.0479609 |
| Control 6h | 54.7713 | 27.4287 | 0.317017 | 0.250571 |
| ABA 6h | 75.0252 | 38.7843 | 0.465165 | 0.308103 |
| ACC 6h | 86.5765 | 29.0905 | 0.578508 | 0.102212 |
| BABA 6h | 135.365 | 52.075 | 0.897807 | 0.315358 |
| Chitin 6h | 100.94 | 22.2987 | 0.769651 | 0.165216 |
| Epi 6h | 296.02 | 28.3951 | 2.20794 | 0.301755 |
| SA 6h | 60.0369 | 9.24079 | 0.502035 | 0.116619 |
| Me-JA 6h | 46.1101 | 31.9838 | 0.210727 | 0.287705 |
Source Transcript PGSC0003DMT400042699 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G29510.1 | +1 | 5e-29 | 100 | 52/98 (53%) | SAUR-like auxin-responsive protein family | chr1:10322683-10323114 FORWARD LENGTH=143 |
