Probe CUST_42426_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42426_PI426222305 | JHI_St_60k_v1 | DMT400052881 | TGCAGTCTCTATGATAGACTTACTCGCATCTGGTGTTTATATCAAACTAATGAATGTAGA |
All Microarray Probes Designed to Gene DMG400020517
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42421_PI426222305 | JHI_St_60k_v1 | DMT400052880 | ACAGTTATGGTGGTATCTCTAGAAGGAAGAATTTGGAAGTCGTCTGGGAAGTACCTCTAA |
| CUST_42426_PI426222305 | JHI_St_60k_v1 | DMT400052881 | TGCAGTCTCTATGATAGACTTACTCGCATCTGGTGTTTATATCAAACTAATGAATGTAGA |
Microarray Signals from CUST_42426_PI426222305
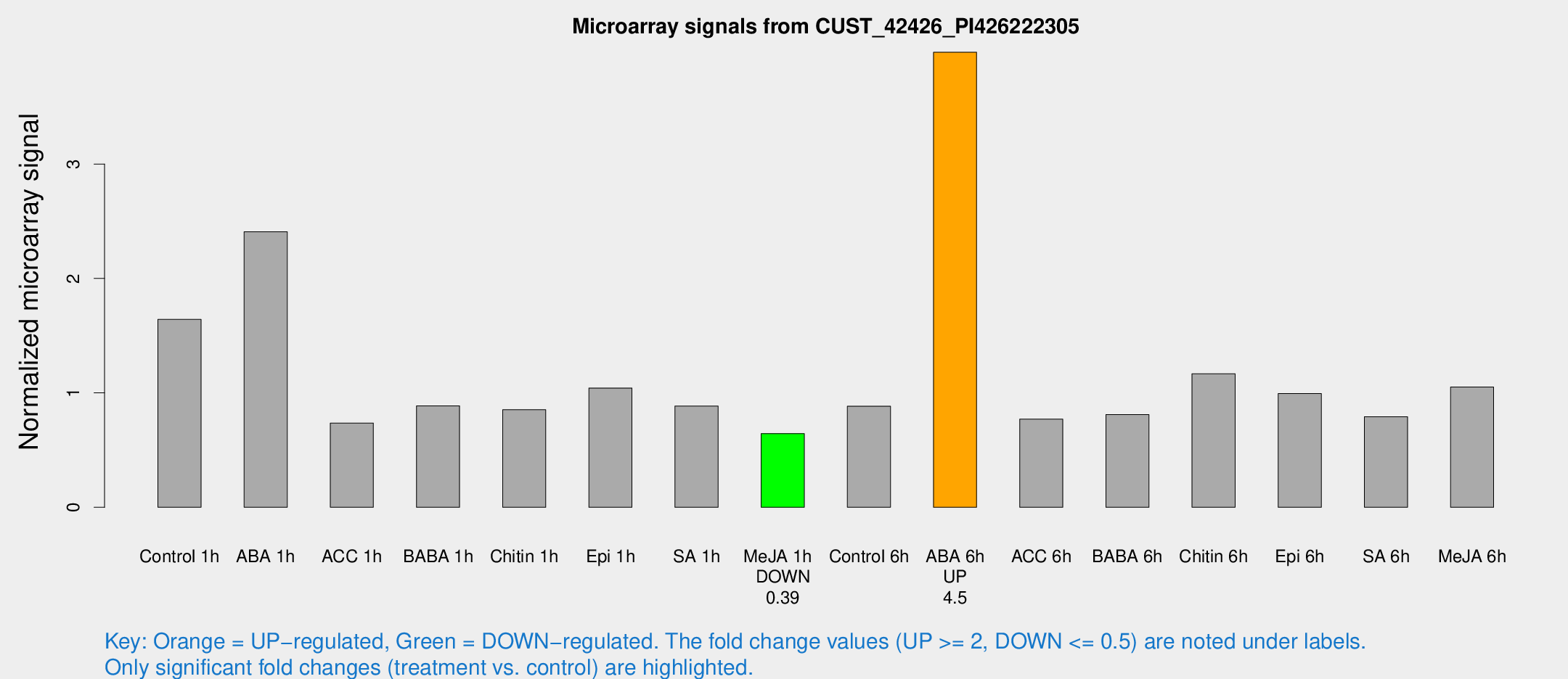
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 155.371 | 14.8106 | 1.64216 | 0.13103 |
| ABA 1h | 199.79 | 11.9633 | 2.40733 | 0.202419 |
| ACC 1h | 81.6647 | 25.3333 | 0.735431 | 0.266572 |
| BABA 1h | 84.7346 | 18.225 | 0.88668 | 0.128822 |
| Chitin 1h | 72.0365 | 5.31404 | 0.852599 | 0.0627372 |
| Epi 1h | 85.5613 | 8.77593 | 1.04234 | 0.100838 |
| SA 1h | 85.6425 | 5.93642 | 0.884816 | 0.11829 |
| Me-JA 1h | 49.6463 | 4.4127 | 0.644382 | 0.0797132 |
| Control 6h | 87.4611 | 17.9315 | 0.883918 | 0.12798 |
| ABA 6h | 409.494 | 68.5856 | 3.97715 | 0.556713 |
| ACC 6h | 84.2069 | 10.3608 | 0.770567 | 0.0581642 |
| BABA 6h | 89.595 | 18.6949 | 0.810635 | 0.168975 |
| Chitin 6h | 116.857 | 7.72174 | 1.16643 | 0.0770425 |
| Epi 6h | 106.278 | 10.8644 | 0.994182 | 0.16848 |
| SA 6h | 76.6401 | 14.5727 | 0.79209 | 0.0763197 |
| Me-JA 6h | 101.626 | 19.1522 | 1.05085 | 0.110412 |
Source Transcript PGSC0003DMT400052881 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G35350.1 | +3 | 0.0 | 782 | 461/810 (57%) | poltergeist like 1 | chr2:14881360-14884116 REVERSE LENGTH=783 |
