Probe CUST_42421_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42421_PI426222305 | JHI_St_60k_v1 | DMT400052880 | ACAGTTATGGTGGTATCTCTAGAAGGAAGAATTTGGAAGTCGTCTGGGAAGTACCTCTAA |
All Microarray Probes Designed to Gene DMG400020517
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42421_PI426222305 | JHI_St_60k_v1 | DMT400052880 | ACAGTTATGGTGGTATCTCTAGAAGGAAGAATTTGGAAGTCGTCTGGGAAGTACCTCTAA |
| CUST_42426_PI426222305 | JHI_St_60k_v1 | DMT400052881 | TGCAGTCTCTATGATAGACTTACTCGCATCTGGTGTTTATATCAAACTAATGAATGTAGA |
Microarray Signals from CUST_42421_PI426222305
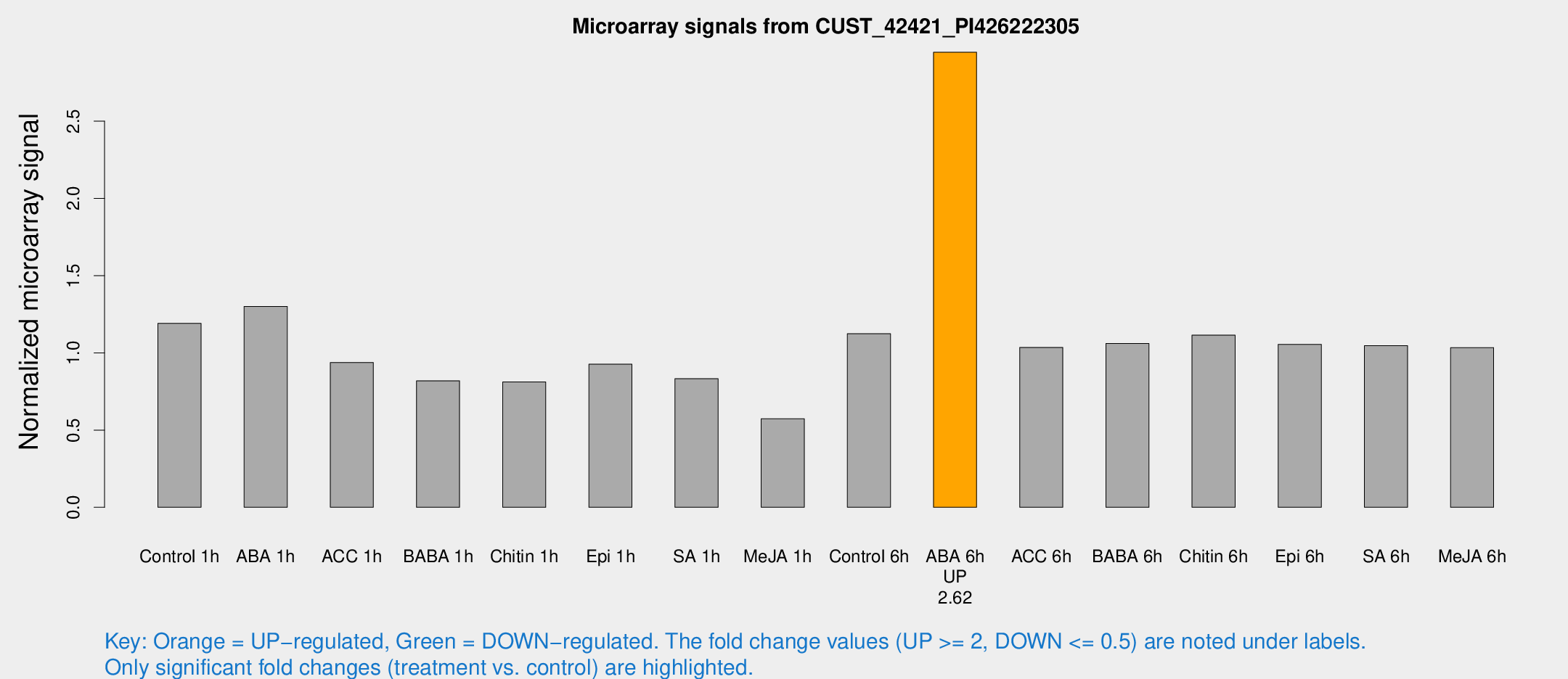
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 289.251 | 43.2484 | 1.19162 | 0.114421 |
| ABA 1h | 274.007 | 18.7568 | 1.30094 | 0.0763234 |
| ACC 1h | 244.704 | 59.5314 | 0.937618 | 0.190137 |
| BABA 1h | 203.442 | 52.2777 | 0.819126 | 0.170188 |
| Chitin 1h | 178.988 | 31.944 | 0.811475 | 0.0925656 |
| Epi 1h | 191.08 | 14.5337 | 0.926638 | 0.0553541 |
| SA 1h | 203.718 | 14.2082 | 0.832728 | 0.0496128 |
| Me-JA 1h | 111.922 | 10.4176 | 0.573287 | 0.102378 |
| Control 6h | 286.067 | 68.8362 | 1.12417 | 0.218408 |
| ABA 6h | 755.052 | 94.4549 | 2.94677 | 0.249308 |
| ACC 6h | 285.036 | 32.1372 | 1.0354 | 0.0611352 |
| BABA 6h | 290.138 | 50.5163 | 1.06071 | 0.164209 |
| Chitin 6h | 284.11 | 27.8071 | 1.11495 | 0.091662 |
| Epi 6h | 284.647 | 28.116 | 1.05449 | 0.0644288 |
| SA 6h | 259.394 | 58.6197 | 1.0463 | 0.144664 |
| Me-JA 6h | 250.628 | 40.1939 | 1.03373 | 0.0847887 |
Source Transcript PGSC0003DMT400052880 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G35350.1 | +1 | 0.0 | 640 | 365/631 (58%) | poltergeist like 1 | chr2:14881360-14884116 REVERSE LENGTH=783 |
