Probe CUST_41262_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41262_PI426222305 | JHI_St_60k_v1 | DMT400077169 | CACCAAAACAGGACAAGTCTGCTTCTAATTATCAGACTAAAGTTACTGATCCAACAGGCG |
All Microarray Probes Designed to Gene DMG400030008
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41262_PI426222305 | JHI_St_60k_v1 | DMT400077169 | CACCAAAACAGGACAAGTCTGCTTCTAATTATCAGACTAAAGTTACTGATCCAACAGGCG |
| CUST_41258_PI426222305 | JHI_St_60k_v1 | DMT400077168 | ACTTGGCAGAGAAATTCAAGCCGCGAGATGAAGATAAAGCACTTTCTGAGGTCATTTCAG |
Microarray Signals from CUST_41262_PI426222305
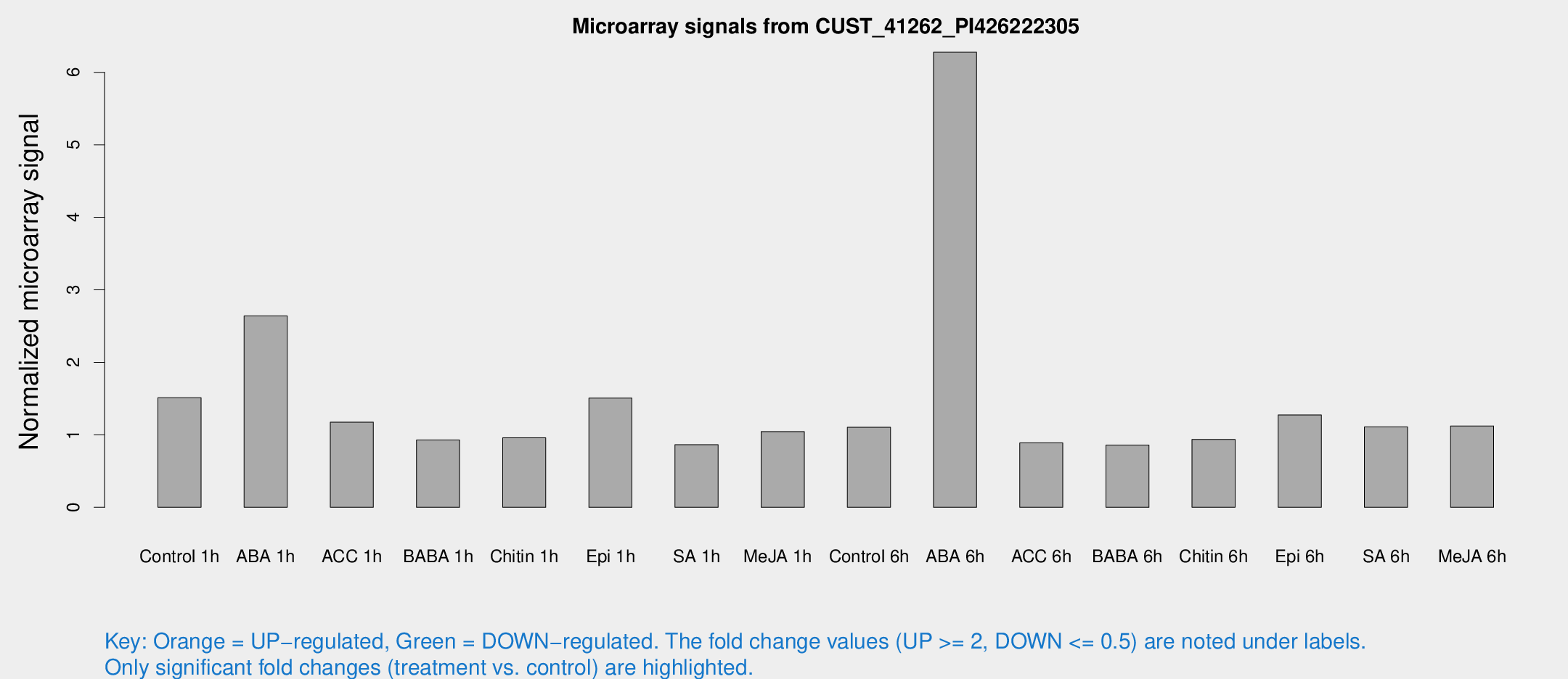
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.9342 | 4.92685 | 1.51175 | 0.588007 |
| ABA 1h | 19.6165 | 7.27144 | 2.64089 | 1.55345 |
| ACC 1h | 8.35507 | 3.47314 | 1.17464 | 0.527704 |
| BABA 1h | 5.97415 | 3.46182 | 0.928976 | 0.537929 |
| Chitin 1h | 5.75971 | 3.21978 | 0.957795 | 0.534933 |
| Epi 1h | 9.1218 | 3.24266 | 1.50593 | 0.603422 |
| SA 1h | 5.94055 | 3.1965 | 0.862826 | 0.468688 |
| Me-JA 1h | 5.70863 | 3.32175 | 1.04476 | 0.605581 |
| Control 6h | 7.75279 | 3.42218 | 1.10468 | 0.541207 |
| ABA 6h | 50.9317 | 15.8592 | 6.27779 | 2.38812 |
| ACC 6h | 6.94482 | 4.11706 | 0.88821 | 0.51452 |
| BABA 6h | 6.42262 | 3.73233 | 0.859014 | 0.498149 |
| Chitin 6h | 6.65211 | 3.77063 | 0.936606 | 0.53128 |
| Epi 6h | 10.6566 | 4.08462 | 1.27335 | 0.602257 |
| SA 6h | 7.60296 | 3.53524 | 1.10807 | 0.555726 |
| Me-JA 6h | 8.0428 | 3.35283 | 1.12123 | 0.541692 |
Source Transcript PGSC0003DMT400077169 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G52300.1 | +3 | 2e-10 | 61 | 48/155 (31%) | CAP160 protein | chr5:21237205-21239404 FORWARD LENGTH=619 |
